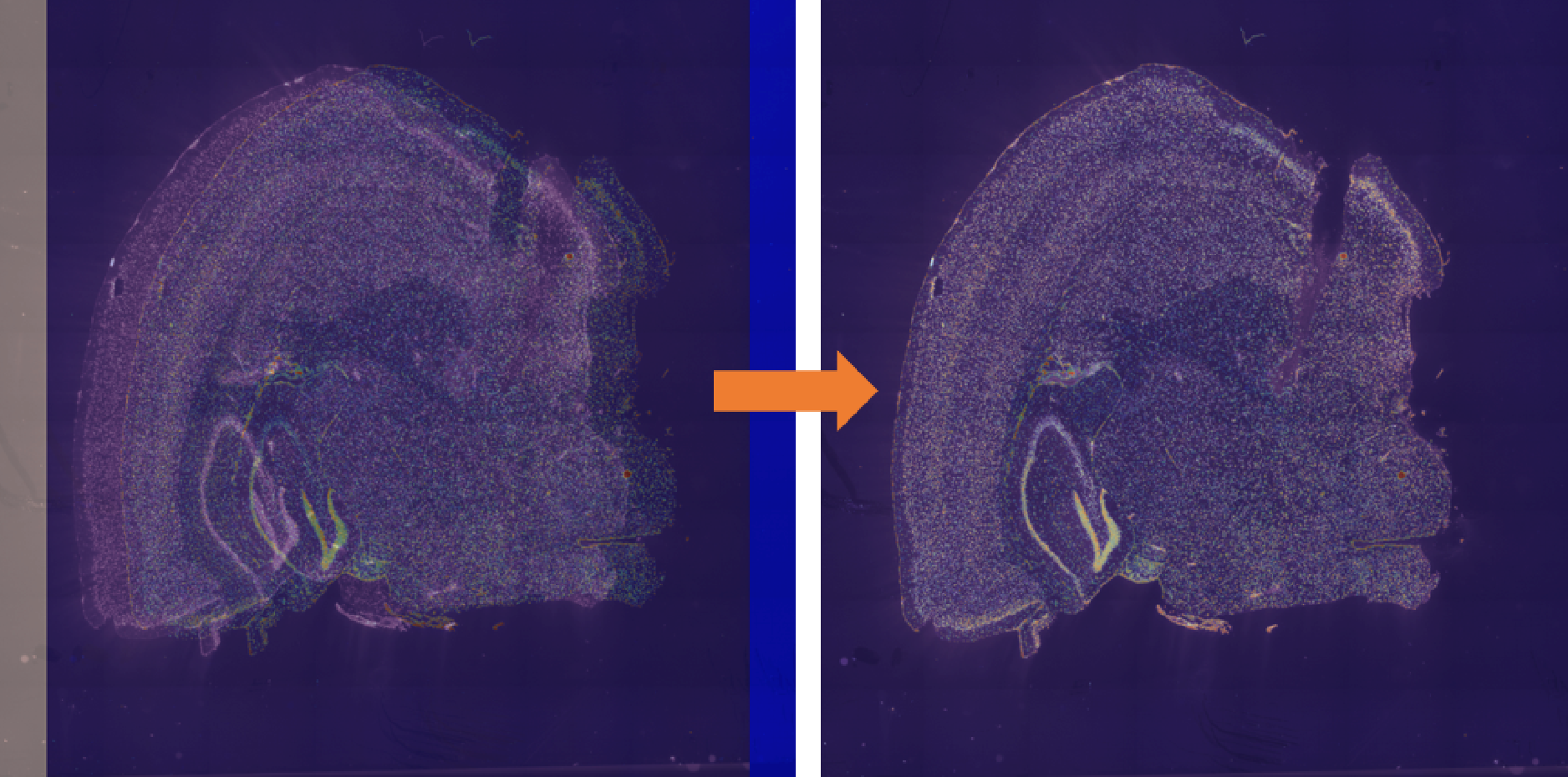
Option 1: use C++ compiled geftools:
Option 2: use Python package - gefpy (e.g. 0.6.1):
Option 3: with installed SAW sif (e.g. v5.1.3):
https://hub.docker.com/repository/docker/stomics/saw
singularity exec SAW_v5.1.3.sif cellCut
Please use Singularity version 3.8 or later
Bash
export HDF5_USE_FILE_LOCKING=FALSE
## gef2gem using geftools
geftools view -i <SN>.gef -o <SN>.gem -s <SN>
# -i input square bin GEF, e.g.SN.raw.gef or SN.gef
# -o output GEM
# -s SN
## gef2gem using gefpy
python
>>> from gefpy.bgef_reader_cy import BgefR
>>> bgef=BgefR(filepath='<SN>.gef',bin_size=200,n_thread=4)
>>> bgef.to_gem('<SN>.bin200.gem')## gef2gem using SAW sif
## export SINGULARITY_BIND="/path/to/input/dir,/path/to/output/dir"
singularity exec SAW_v5.1.3.sif cellCut view -i <SN>.gef -o <SN>.gem -s <SN>
## cgef2cgem
geftools view -i <SN>.cellbin.gef -o <SN>.cellbin.gem -d <SN>.raw.gef -s <SN>
# -i input cellbin GEF
# -o output cellbin GEM
# -d input square bin GEF, e.g. SN.raw.gef or SN.gef
# -s SN
## gem2gef
geftools bgef -i <SN>.gem -o <SN>.gef -b 1,20,50 -O Transcriptomics
# -i input square bin GEM
# -o output square bin GEF
# -b bin sizes seqarate by comma, default: 1,10,20,50,100,200,500
# -O omics name