Q
How to limit contamination between cutting different samples?
A
It is recommended to replace the blades when cutting different samples, and to clean the sectioning area before the next sample. It is best to clean the sectioning area and brushes with ethanol before and after sectioning to avoid cross-contamination of samples.
Q
What is the thickness of the tissue section for Stereo-seq Transcriptomics Set?
A
The general sectioning thickness is 10 μm, and the thickness can be adjusted according to the research purpose.
Q
What are the requirements of Stereo-seq Transcriptomics Set for imaging systems?
A
Please refer to Microscope Assessment Guideline for specific requirements and evaluation of microscope.
Q
Is there a standard cDNA yield range?
A
cDNA yield depends on many factors, such as tissue type, cell type, tissue size and tissue RNA quality. There is no standard cDNA yield range.
Q
What are the sizes of the spots on the Stereo-seq Chip T (1cm*1cm)?
A
The spots are 220 nm in diameter.
Q
Why is my cDNA yield too low?
A
This could be caused by low cell numbers,RNA degradation in the tissue or many other factors induced during the experimental procedures. Also, it may be caused by sample loss during experimental procedures or insufficient cDNA release during cDNA release step.
Q
What is the center-to-center distance between two spots?
A
The center-to-center distance between two spots is 500 nm.
Q
How should unused chips be stored?
A
Store it in TE buffer at 4℃.
Q
What is the FF library structure and recommended sequencing strategy?
A
Library structure is indicated in the figure below:
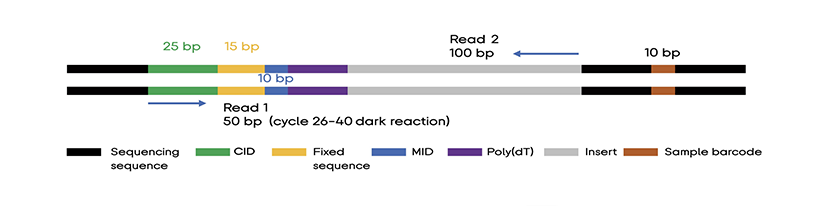
Use the following configuration to perform paired-ended sequencing run:
Read 1: 50 cycles (25 cycles for barcode + 15 dark cycles for fixed sequence +10 cycles for MID),
Read 2: 100 cycles
Sample barcode:10 cycles
The library construct is compatible with our DNBSEQ platform. Users are free to try out other platforms.
Q
Why are there track lines showing on the Stereo-seq Chip T (1cm*1cm) after ssDNA/DAPI staining?
A
After ssDNA/DAPI staining, the fluorescent signal contrast between the capturing probe loaded area and unloaded area allows visualization of the track lines.