All
Products
Resources
News
FAQ
Search
10/02/2026
Within the field of spatial biology, the scale of observation directly shapes the questions a researcher can ask. A fundamental choice exists between methods that analyze small, selected regions and those capable of capturing an entire tissue section. This discussion focuses on the operational and scientific distinctions between standard small-field methods and emerging large-area transcriptomics approaches. At STOmics, we have focused our development on enabling comprehensive large stereo-seq transcriptomics to address the limitations of restricted field-of-view analysis.
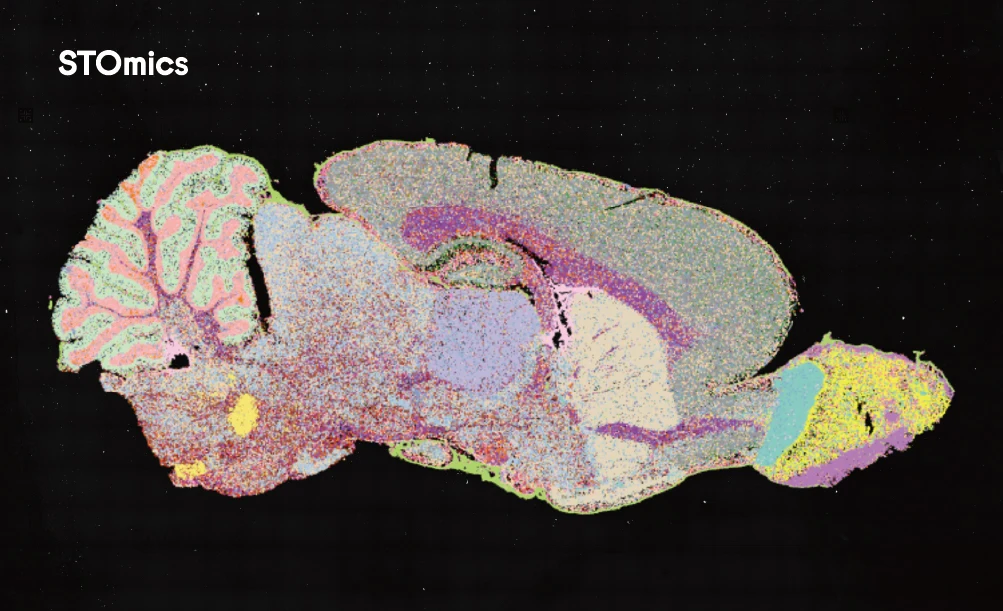
Traditional small-field spatial transcriptomics methods often analyze areas measuring only a few square millimeters. This restricted view necessitates a difficult choice: researchers must either select a few tiny, representative regions or stitch together numerous individual captures to approximate a larger area. The first approach risks missing critical biology occurring outside the chosen fields. The second method, image stitching, can introduce technical artifacts and requires complex, often imperfect alignment of data seams. These limitations mean that studying an entire organ cross-section, a complete tumor biopsy with its heterogeneous microenvironment, or developmental structures at a true tissue scale becomes a major logistical and analytical challenge. The view through a keyhole, while detailed, can never reveal the layout of the entire room.
In contrast, large-area transcriptomics is defined by its capacity to capture data from centimeter-sized fields of view in a single, seamless experiment. This approach, exemplified by our large stereo seq transcriptomics using chips up to 2cm x 3cm, removes the need for pre-selection or stitching. An entire tissue section can be placed on one chip, preserving its full anatomical context. Every cell, from the core of a lesion to the surrounding stroma and healthy margins, is captured simultaneously on a unified spatial coordinate system. This ensures that the complex spatial relationships and gradients across the entire sample are retained intact. The move to a large canvas allows the biology itself, not the instrument's limitations, to define the boundaries of the experiment.
The practical impact of this scale is significant. In pathology, a whole biopsy can be analyzed at once, enabling the unbiased correlation of molecular profiles with all recognized histological regions. For developmental biology, researchers can map gene expression patterns across an entire embryo section, observing organizing centers and morphological gradients without interruption. In neuroscience, architectural studies of brain layers and regions benefit from an unimpeded view. This comprehensive capture is the core of large-area transcriptomics. The large stereo-seq transcriptomics platform from STOmics delivers this by combining subcellular resolution with a centimeter-scale field of view, providing a "tissue-to-data" solution that captures the whole transcriptome across complete sections.
The comparison between methods ultimately centers on scope and context. While small-field analyses provide valuable local data, they can inadvertently obscure the bigger picture. The progression toward large-area transcriptomics represents a shift towards preserving and analyzing biological systems in their complete architectural state. By offering a platform for large stereo-seq transcriptomics on chips designed for entire tissue sections, we at STOmics provide researchers with the tools to move from analyzing snapshots to studying the full landscape, ensuring that critical spatial relationships are never cropped out of the data.