Tumors are highly heterogeneous tissues characterized by intricate spatial architecture, wherein cancer cells, immune cells, and stromal cells collectively constitute distinct and dynamic microenvironments across the tumor landscape(Pattabiram et al., 2025). Understanding spatial heterogeneity is critical for uncovering how tumors work and why treatments succeed or fail, but traditional transcriptomic methods cannot capture this complexity. Stereo-seq enables entire tissue sections to be studied at single-cell resolution, revealing how spatial cell interactions shape tumor behavior.
Limitations of Traditional Methods
Traditional transcriptomic approaches have limitations:
Bulk RNA-seq averages signals, masking cellular heterogeneity (Li et al., 2021).
Single-cell RNA-seq preserves cell identity but loses spatial context due to tissue dissociation (Li et al., 2021).
Other spatial transcriptomics trade off resolution for coverage, limiting the ability to capture both fine cellular details and large-scale structures like invasive margins or gradients (Ahmed et al., 2022).
Thus, existing methods struggle to reveal how spatially organized microenvironments drive tumor progression.
Stereo-seq enables whole-section, single-cell spatial transcriptomics at scale
Stereo-seq enables researchers to map cellular heterogeneity and spatial organization across entire tumor sections without sacrificing resolution. It supports precise cell type annotation and direct visualization of cell–cell interactions in both formalin-fixed paraffin-embedded (FFPE) and fresh frozen (FF) samples.
This performance is achieved using ultra-high-density DNA Nanoballs (DNBs) arrays with 500 nm spacing, which preserve native tissue architecture while capturing the whole transcriptome (Figure 1). With chip sizes scalable up to 13 × 13 cm, Stereo-seq enables spatial profiling across large tissue areas and entire tumor sections (Figure 2). This scalability is achieved without compromising single-cell resolution, enabling analysis from local microenvironments to whole-section contexts.
 arranged at single-cell resolution, enabling mRNA from individual cells to be detected with hundreds of DNBs, each carrying a unique Coordinate ID (CID)..png)
Figure 1. Stereo-seq captures DNA Nanoballs (DNBs) arranged at single-cell resolution, enabling mRNA from individual cells to be detected with hundreds of DNBs, each carrying a unique Coordinate ID (CID).
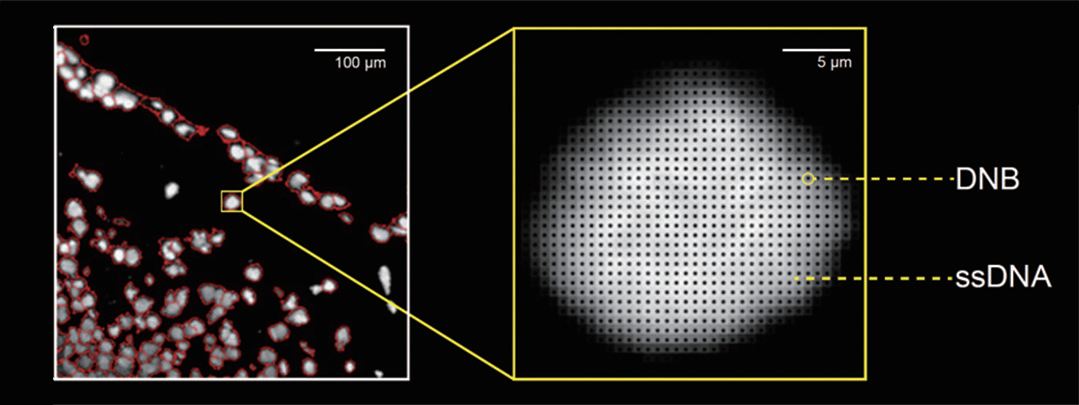
Figure 2. Demonstration of Stereo-seq chips of different sizes.
Case Studies: Stereo-seq in Action
To illustrate the power of Stereo-seq in real-world applications, we highlight two representative case studies: Total Neoadjuvant Therapy (TNT) Response in Rectal Cancer and Immune Architecture in Cervical Squamous Cell Carcinoma (CSCC).
Spatial Insights into TNT Response in Rectal Cancer
Using Stereo-seq Transcriptomics FF Solution, Gao et al. (2025) visualized how immune cells are organized across entire tumor sections and how these spatial patterns evolve during treatment. Researchers observed that strong responders consistently exhibited ACKR1⁺ endothelial cells (ECs) closely colocalized with IFNG⁺ CD8⁺ T cells, forming immune–vascular niches that sustain T-cell recruitment and enhance tumor killing (Figure 3). Access to this spatial context uncovers patterns that would otherwise remain invisible, providing critical insights into the immune mechanisms behind TNT efficacy.
..jpeg)
Figure 3. Spatial transcriptomics shows ACKR1⁺ endothelial cells adjacent to IFNG⁺/GZMH⁺ CD8⁺ T cells in TNT-treated tumors, suggesting immune–vascular niche formation. Source: Gao et al., Cancer Cell (2025).
Spatial Mapping of Immune Architecture in Cervical Cancer
In a study by Fan et al. (2023), researchers used the Stereo-seq Transcriptomics FF Solution to profile entire CSCC tumor sections at high spatial resolution, enabling a comprehensive view of tumor–immune architecture across the whole tissue.This spatial analysis allowed direct comparison of tumor regions dominated by the immune-resistant MP6 state and the immune-infiltrated MP7 state within their native context. As a result, tumor ecosystems associated with immune exclusion (MP6-dominant) and immune infiltration (MP7-dominant) were clearly distinguished, providing direct spatial evidence for differential immunotherapy responses (Figure 4).
..png)
Figure 4. Stereo-seq revealed that MP6 and MP7 states occupy spatially distinct tumor regions. Source: Fan et al., Nature Genetics (2023).
STOmics Product Matrix: Scalable Solutions Across Spatial Scales
Following our product case studies, STOmics uses core Stereo-seq technology to deliver spatial transcriptomics solutions. The product matrix (Table 1) shows available options and typical applications. All products share the same ultra-high-density DNA Nanoballs (DNBs) array, ensuring consistent single-cell resolution and transcriptome depth.
Table 1. STOmics product matrix
Looking ahead: The future of spatial cancer research
Technologies like Stereo-seq are transforming how we study tumors, from identifying new cellular interactions to understanding how microenvironments affect treatment outcomes. These insights will help design more precise therapies and accelerate translational research.
Need tailored spatial transcriptomics solutions or guidance on experimental design? Feel free to Contact Us or reach out directly at info_global@stomics.tech. Our team is here to help you achieve your research goals.
