Cervical Squamous Cell Carcinoma (CSCC) is the most common form of cervical cancer and represents a significant area of research in oncology. CSCC is often diagnosed at advanced stages, where metastasis and recurrence become significant concerns. Recent studies have focused on understanding the molecular mechanisms underlying tumor progression, immune evasion, and treatment resistance in order to identify more effective therapeutic strategies. A key area of focus is understanding the role of the tumor immune microenvironment (TIME), including immune cell interactions and immune suppression mechanisms, in shaping the course of the disease (Cohen et al., 2019).
Current Challenges in Clinical Treatment
CSCC presents numerous clinical challenges, one of the most critical being its poor response to immune checkpoint blockade (ICB) therapies, including PD-1/PD-L1 inhibitors, which have shown limited efficacy in CSCC. The objective response rate (ORR) reported in PD-L1–positive CSCC patients receiving pembrolizumab is only around 17%, highlighting the difficulty in overcoming immune resistance. Immune exclusion is a major factor contributing to the limited efficacy of immunotherapy in CSCC, where immune cells accumulate around the tumor but fail to effectively penetrate the tumor core, resulting in immune escape and treatment resistance (Shieh et al., 2021). Increasing evidence suggests that this phenomenon is closely associated with the heterogeneous expression of tumor- and stroma-related signaling pathways within the tumor microenvironment, which can create localized immunosuppressive niches that restrict immune cell infiltration. Understanding how these heterogeneous cellular states and signaling programs are spatially organized within tumors has therefore become critical for uncovering the mechanisms of immune exclusion and developing more effective therapeutic strategies.
How Stereo-seq Enhances Spatial Resolution Mapping
Fan et al. (2023) analyzed 31 treatment-naïve CSCC samples using a combination of single-cell RNA sequencing (scRNA-seq) and Stereo-seq Transcriptomics Solution to map the heterogeneity of CSCC tumors at an unprecedented resolution. The study applied Non-negative Matrix Factorization (NMF) to scRNA-seq data to identify seven distinct transcriptional meta-programs (MP1–MP7) in CSCC cells, and these states were then spatially mapped using Stereo-seq, a high-resolution spatial transcriptomics platform developed by STOmics (Figure 1).
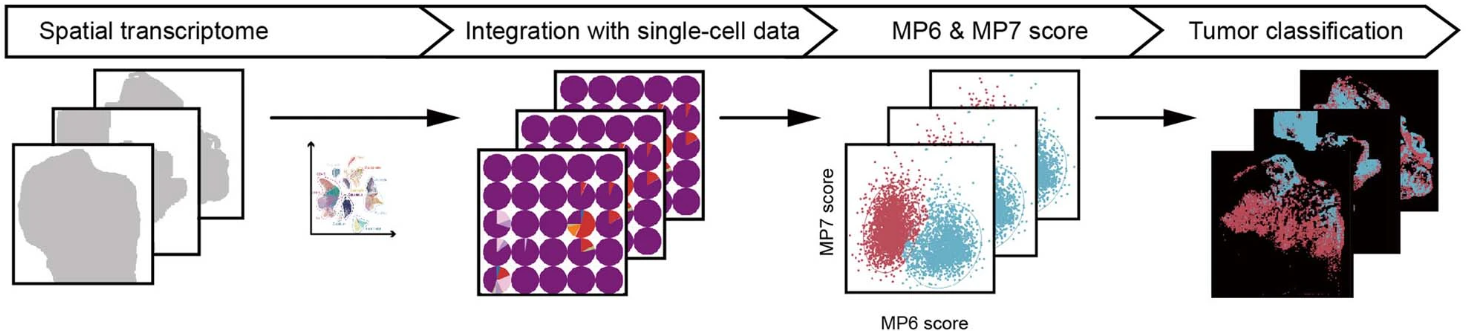
Figure 1. Workflow for identifying MP6 and MP7 tumor spots by integrating scRNA-Seq and Stereo-seq data.
The study classified malignant CSCC cells into multiple meta-programs (MPs), particularly focusing on MP6 and MP7, which represented immune-excluded and immune-infiltrated states, respectively (Figure 2). By applying ligand-receptor interaction and spatial proteomics to the research, it was found that MP6 tumors were associated with immune exclusion, displaying features of keratinization and high expression of TGFβ signaling, which led to the recruitment of immunosuppressive cancer-associated fibroblasts (CAFs). On the other hand, MP7 tumors displayed immune infiltration, with enriched immune signatures associated with T cell activation and interferon signaling.
 Use Stereo-seq to detect the spatial distribution of tumor spots in several slices. (b) Different MP6 and MP7 regions are presented by mapping onto a Stereo-seq map..png)
Figure 2. (a) Use Stereo-seq to detect the spatial distribution of tumor spots in several slices. (b) Different MP6 and MP7 regions are presented by mapping onto a Stereo-seq map.
Why do we need Stereo-seq?
Traditional scRNA-seq has helped researchers understand the molecular makeup of tumors, identifying different cellular states within a tumor (Ahmed et al., 2022). However, scRNA-seq does not retain spatial context, making it difficult to map the interaction between tumor cells and immune cells in the tumor microenvironment. This lack of spatial information limits our ability to understand how specific cell states (e.g., immune exclusion) relate to immune cell infiltration patterns, a gap that prevents the development of targeted therapies to overcome immune evasion in tumors like CSCC. This limitation highlights the need for spatial transcriptomics technologies, which provide the resolution needed to link molecular data to tissue architecture.
The breakthrough of this research was made possible by the Stereo-seq Transcriptomics Solution, which bridges this gap by restoring the critical spatial coordinates to transcriptomic data. While other spatial transcriptomics platforms typically operate at micron-level resolution with limited capture areas, Stereo-seq stands out with its 500nm sub-cellular resolution compared to 55 µm (Visium) or single-molecule FISH-based approaches. This unparalleled precision allowed researchers to move beyond simple cell lists to visualize the exact structure of the immune area. By integrating high-resolution spatial mapping with a centimeter-level field of view, Stereo-seq provides distinct advantages for studies requiring both high resolution and large tissue coverage, enabling a more accurate representation of tumor biology and identifying the precise molecular barriers to immunotherapy.
Without Stereo-seq, researchers would only have been able to observe the general upregulation of FABP5 and TGFβ pathways within the tumor bulk, without understanding the specific spatial context. The precise spatial mapping enabled by Stereo-seq revealed how malignant cells (specifically those in the MP6 program) interact with CAFs to form a dense, fibrotic structure that prevents immune cell infiltration. By capturing the spatial co-occurrence of transcripts at a sub-cellular level, the study showed that the FABP5-TGFβ signaling axis plays a central role in immune exclusion. This high-resolution mapping led to the development of a spatial niche score, a more accurate metric than traditional bulk or single-cell signatures, as it accounts for the physical and chemical barriers that block T-cell infiltration (Figure 3).
Figure 3. Spatial structures of MP6 and MP7 regions. MP7 shows spatial co-localization of malignant regions with immune inflammatory genes, while the absence of immune signaling in MP6 reveals its immune exclusion characteristics.
STOmics Product Matrix: Scalable Solutions Across Spatial Scales
The CSCC study utilized the Stereo-seq Transcriptomics Solution. STOmics offers a comprehensive suite of spatial transcriptomics solutions for high-resolution analysis of gene expression within intact tissue architecture. All solutions are based on the same ultra-high-density DNA Nanoball (DNB) array, ensuring consistent single-cell–level resolution and deep transcriptome coverage across applications. The product matrix below summarizes the available Stereo-seq solutions and their typical applications.
Table 1. STOmics product matrix
The Future is Spatial
As we look toward the future of precision medicine, the ability to resolve the spatial architecture of disease will be the defining factor in therapeutic discovery. Stereo-seq is not just a sequencing tool; it is a spatial navigation system for the next generation of oncology. To the PI or professor aiming to transcend descriptive single-cell data, Stereo-seq provides the resolution and scale to secure high-impact, clinically relevant publications.
Explore how Stereo-seq can advance your spatial biology research →
Contact Us or email us at info_global@stomics.tech for tailored experimental design and support in achieving your research goals.
