Regeneration is an ancient biological trait that has been inconsistently preserved across evolution. Many animals, such as salamanders and fish, are capable of regenerating complex tissues after injury, including limbs and internal organs (Avila-Martinez et al., 2023). In contrast, most mammals respond to injury primarily through fibrotic repair, forming scar tissue that restores structural integrity but often results in permanent loss of function (Bely & Nyberg, 2010). This striking difference across species has raised a fundamental question in regenerative biology: why is regenerative capacity so limited in mammals?
Current Gap Leads to Problems
The core of regeneration is the formation of a blastema, a region of progenitor cells that drive tissue regrowth. Without spatial data, it’s hard to distinguish between fibroblasts that cause scarring and those with regenerative potential, as they often share similar markers but occupy different tissue regions (Tower et al., 2022). Additionally, comparing mammals with species like salamanders is challenging due to their vast genetic differences. A method to compare regeneration within mammals was needed to identify the evolutionary “lesions” that inhibit regeneration.
Mapping Regenerative Mechanisms with Stereo-seq
To investigate the molecular mechanisms behind mammalian regenerative limitations, the researchers combined Stereo-seq, scRNA-seq, and functional validation, with their findings published in the January 2025 issue of the prestigious journal Science (Lin et al., 2025).They analyzed injury sites in regenerative (rabbits) and non-regenerative species (macaques, mice, and rats) by collecting ear pinna samples at multiple time points post-injury.
Using the STOmics' Stereo-seq Transcriptomics Solution, they mapped the spatial distribution of cell populations and identified wound-induced fibroblasts (WIFs), which were central to fibrosis (Figure 1). While scRNA-seq identified fibroblast subtypes, it couldn’t determine their spatial organization. Integrating spatial data from Stereo-seq allowed the researchers to map these subtypes to specific injury regions, revealing gene expression patterns related to fibrosis.
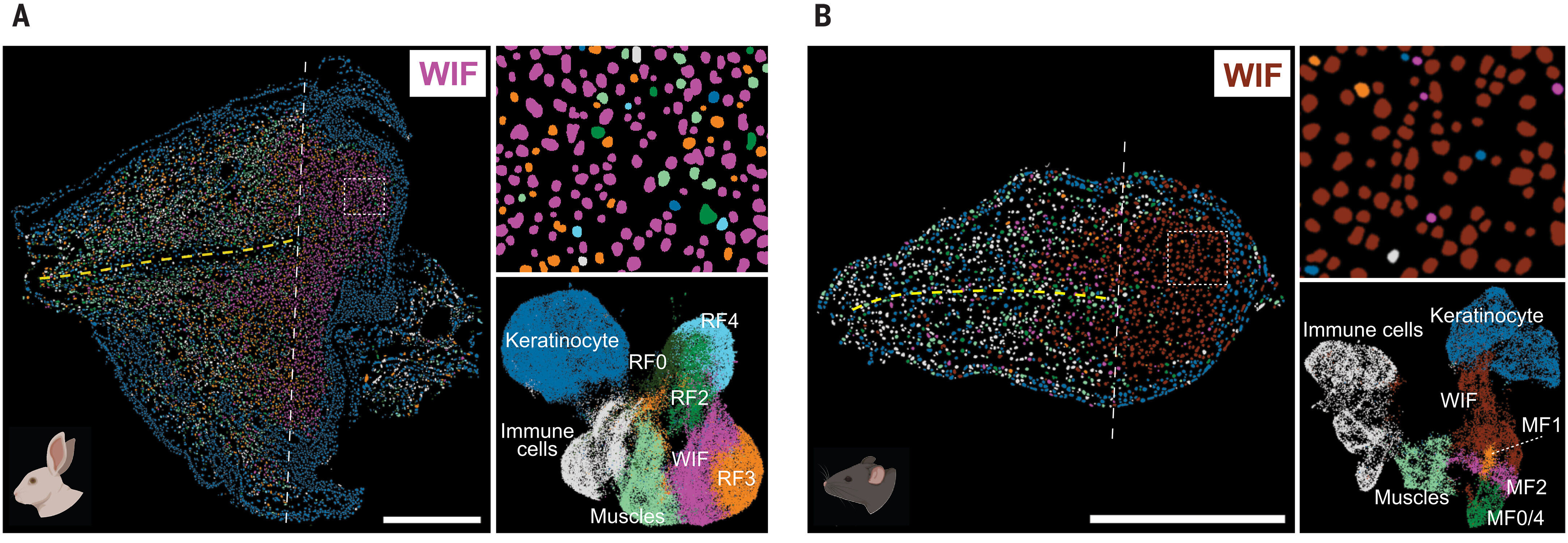
Figure 1. Mapping the Spatial Architecture of Fibrosis. High-resolution spatial transcriptomics via STOmics Stereo-seq (500 nm resolution) compares the regenerative rabbit ear pinna (A) alongside the non-regenerative mouse model (B). This spatial context allows for the precise visualization of distinct wound-induced fibroblast (WIF) subclusters previously indistinguishable by traditional scRNA-seq.
The study identified a deficiency in Retinoic Acid (RA) signaling, linked to the downregulation of Aldh1a2, as a key factor in the failure of regeneration. Functional validation showed that activating the RA/Aldh1a2 pathway shifted repair from fibrosis to regeneration, demonstrating Stereo-seq’s power in mapping spatial molecular interactions and providing insights into the regenerative process (Figure 2).
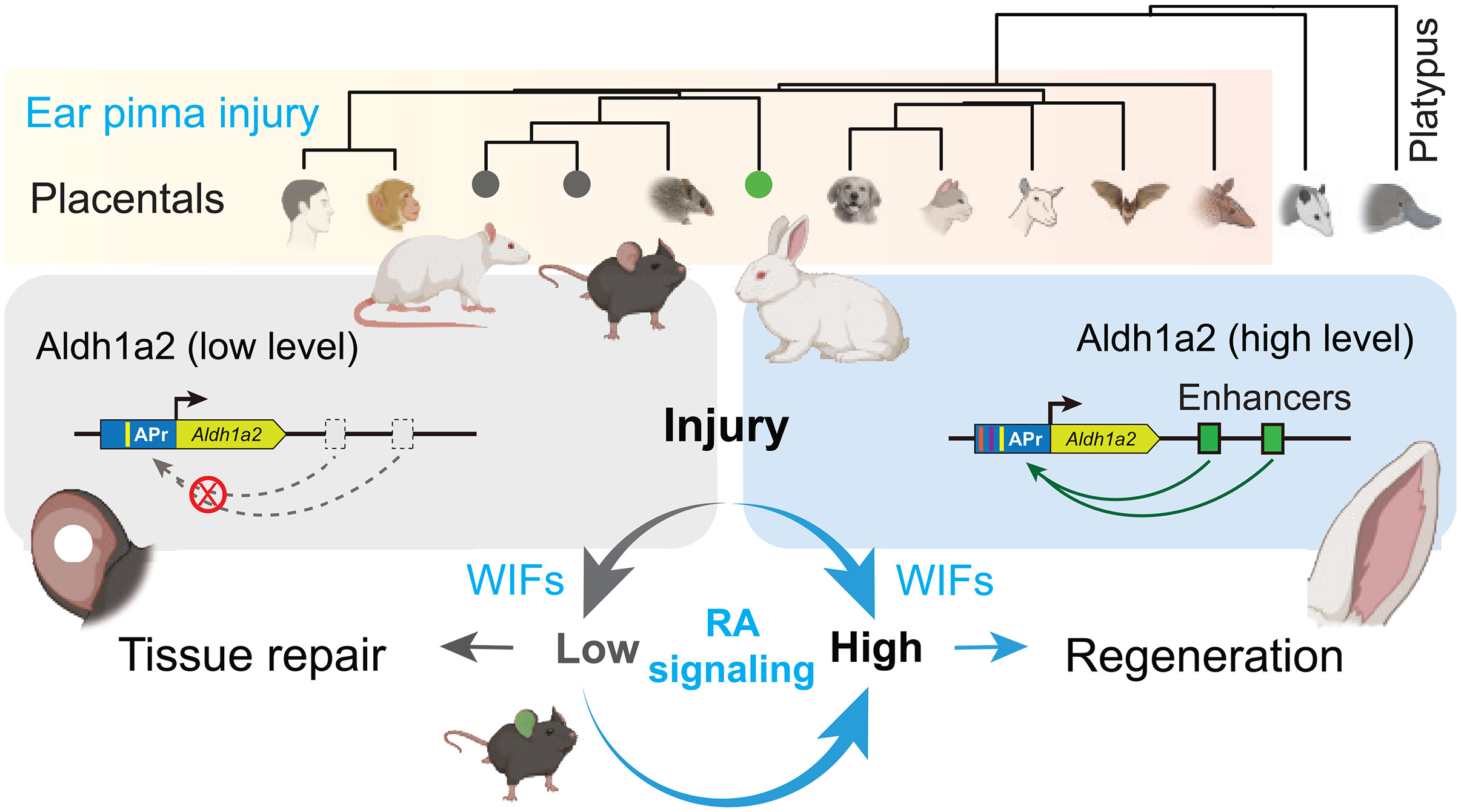
Figure 2. Rethinking Mammalian Regenerative Evolution. A mechanistic schematic enabled by STOmics spatial mapping, revealing that mammalian regeneration is not genetically lost, but spatially suppressed. It illustrates how Aldh1a2-driven Retinoic Acid (RA) signaling actively shifts the tissue repair trajectory from permanent fibrosis (mice) toward complete regeneration (rabbits).
The Stereo-seq Advantage: From “Cellular Soup” to Spatial Architecture
The core of this discovery lies in the ability to visualize the “Regenerative Pocket”, a critical aspect that previous non-spatial methods could not capture. Stereo-seq overcame the bottleneck by providing a high-resolution, spatially aware approach that allowed researchers to pinpoint key regenerative processes.
Without Stereo-seq, traditional methods would only highlight a general increase in fibroblast activity, without revealing the specific locations where this activity occurs—whether at the wound edge, in cartilage, or scattered throughout the tissue. This lack of spatial precision made it impossible to identify the critical signaling center necessary for blastema formation, hindering progress in regenerative studies.
Using STOmics' Stereo-seq technology, researchers could now visualize distinct fibroblast subtypes and their spatial organization (Figure 3). The team identified that in rabbits, a specific subset of Aldh1a2 fibroblasts migrates to the injury site to create a regenerative environment, a detail that would have been missed without spatial mapping (Figure 4). Furthermore, Stereo-seq revealed precise gradients of RA signaling, showing how, in mice, the Aldh1a2 pathway is spatially silenced at the wound site, triggering fibrotic repair instead of tissue remodeling.
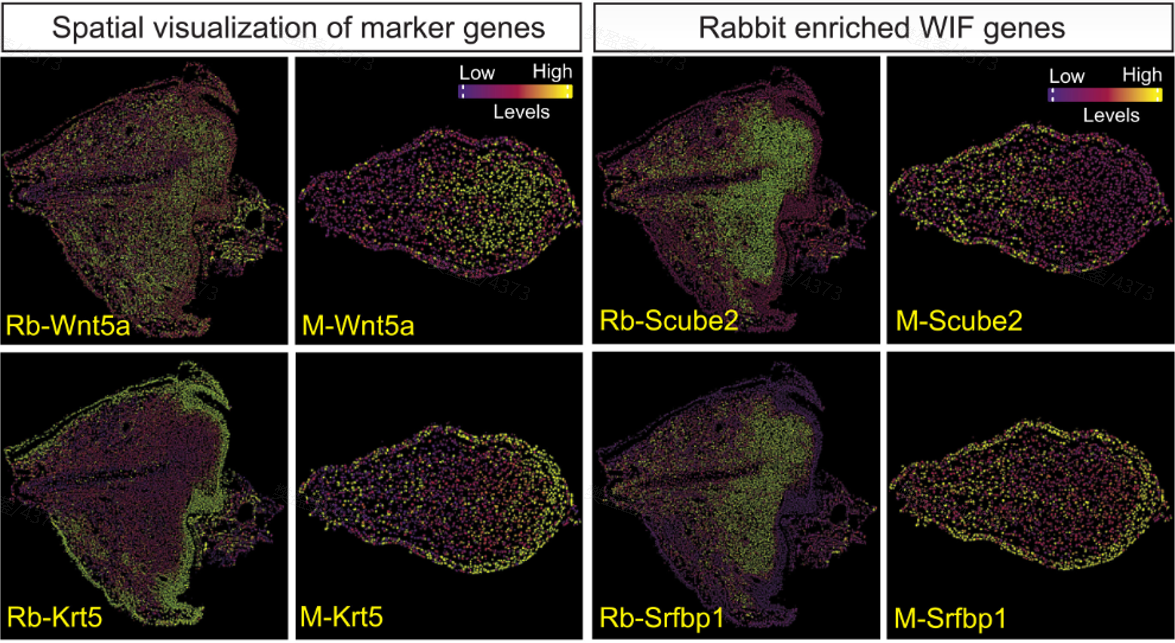
Figure 3. Visualizing the "Regenerative Pocket". Spatial visualization of key blastema-related marker genes (Wnt5a, Krt5, Scube2, and Srfbp1) using Stereo-seq technology. The distinct expression gradients highlight the divergent microenvironments between the regenerative rabbit tissue (left) and the fibrotic mouse model (right), a crucial insight unattainable without true nanoscale spatial mapping.
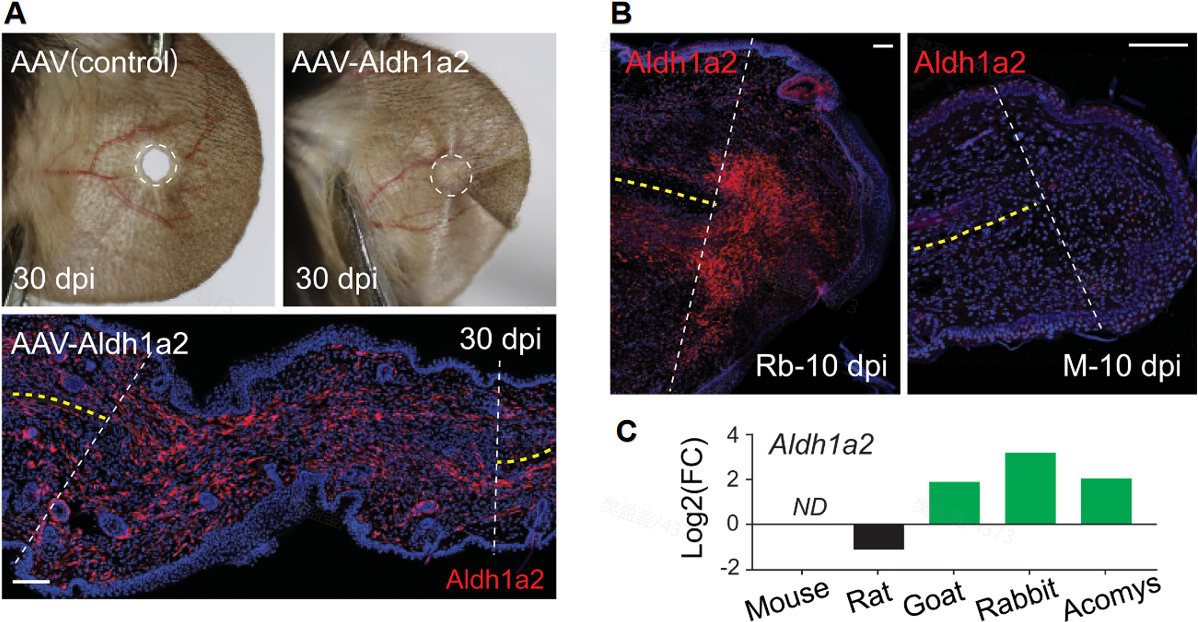
Figure 4. Functional Validation of the Spatial Switch. (A) Aldh1a2 overexpression via AAV successfully restores ear pinna regeneration in mice at 30 days post-injury (dpi). Supported by spatial transcriptomics data, (B) Aldh1a2 is shown to be highly activated in the rabbit ear pinna at 10 dpi, yet absent in the mouse model. (C) Consequently, Aldh1a2 is upregulated exclusively within the regenerating microenvironment, proving that manipulating spatial signaling can reactivate latent healing potential.
The ability to map these spatial gene expressions was pivotal in demonstrating that mammalian regeneration is not lost, but rather spatially suppressed. Stereo-seq provided the spatial roadmap to flip the switch from scarring to healing, proving that the regenerative blueprint is still embedded within the mammalian genome.
STOmics Update: A Total Solution Matrix for Spatial Omics
Building on discoveries like this spatial insight into mammalian regeneration in Science, STOmics provides a comprehensive suite of spatiotemporal omics solutions designed to unlock the full potential of your next research project. Table 1 summarizes the key technical specifications of each solution and highlights their representative applications in spatial biology research.
Table 1. STOmics product matrix
A New Era of Spatial Technology
The success of this study demonstrates that Stereo-seq is more than just a mapping tool—it is a discovery engine for therapeutic intervention, which will allow clinical researchers to “wake up” pathological samples and study complex organ-level regeneration in humans.
For PIs and Lead Researchers, Stereo-seq offers the ability to bridge the gap between evolutionary biology and clinical application, ensuring that no cellular interaction goes unnoticed. For tailored experimental design and support in achieving your research goals, feel free to Contact Us at info_global@stomics.tech.
FAQs
Why is spatial transcriptomics important for tissue regeneration?
What makes Stereo-seq different from other spatial transcriptomics technologies?
Which Stereo-seq solution should I choose?
Stereo-seq Transcriptomics Solution: The golden standard for fresh frozen (FF) tissues to capture high-resolution spatial gene expression.
Stereo-seq OMNI Solution: Optimized for FFPE samples; the ideal choice for unlocking insights from clinical and archived tissue banks.
Stereo-seq Large Chip Designs: Designed for whole-organ mounting (e.g., monkey brains), offering extensive centimeter-scale area coverage.
- Stereo-CITE Solution: The premier choice for spatial multi-omics, enabling simultaneous spatial transcriptomics and ultra-high-plex protein detection.
Spatial transcriptomics allows precise mapping of gene expression within intact tissues. Unlike traditional bulk RNA-seq, which creates a "cellular soup," technologies like STOmics' Stereo-seq preserve the spatial architecture, revealing critical microenvironments—such as the elusive 'Regenerative Pocket'—where regeneration initiates.
STOmics' Stereo-seq sets itself apart with its industry-leading 500 nm true nanoscale resolution and uniquely scalable capture area (centimeter-scale up to 13×13 cm). This allows researchers to capture whole-organ landscapes and sub-cellular details seamlessly in a single chip, making it arguably the most powerful tool for complex regenerative studiesand whole-organ atlas projects.
