Gastric cancer remains a leading cause of global cancer mortality, characterized by significant geographic disparities in incidence. The development of intestinal-type gastric cancer typically follows the Correa cascade—a progressive transformation from chronic inflammation through intestinal metaplasia (IM) and dysplasia to carcinoma (Huang et al., 2025). As a critical premalignant phase, IM provides a unique window into early tumor evolution. While Helicobacter pylori (Hp) and host genetics are primary drivers, emerging evidence indicates that the localized immune landscape and microbial communities also direct disease progression (Li et al., 2025). Deciphering these complex spatial interactions is vital for enhancing early detection and precision preventive strategies.
Technical Limitations in Gastric Cancer Research
Genomic sequencing has identified key drivers in gastric cancer, including BRAF, KRAS, ARID1A, and the SBS17 mutational signature enriched in IM (Huang et al., 2023). However, conventional approaches face a critical limitation: the loss of spatial context.
Bulk and single-cell sequencing require tissue dissociation, destroying architectural relationships between epithelial cells, immune infiltration, and microbes like Streptococcus. This makes it impossible to determine whether specific mutations localize to areas of high microbial density or particular immune niches (Zhong et al., 2026). Additionally, valuable FFPE clinical specimens suffer from RNA degradation, limiting transcriptomic analysis. Most technologies also cannot simultaneously detect human and bacterial transcripts in situ. These gaps prevent understanding how genetic alterations, immune responses, and microbial colonization collectively drive progression from IM to cancer.
Integrated Genomic and Spatial Analysis with Stereo-seq
To investigate the molecular evolution of IM, a novel study published in Cancer Discovery on March 2, 2026, implemented a multi-layered experimental strategy integrating genomic sequencing, epigenomic profiling, and spatial transcriptomics across more than 1,500 IM samples from different countries (Huang et al., 2026). High-depth targeted DNA sequencing with an average coverage exceeding 1,100× identified over 2,100 driver mutations across 47 significantly mutated genes. Whole-genome sequencing further revealed distinct mutational processes such as the SBS17 signature, which appears specifically enriched in IM compared with normal gastric epithelium. Transcriptomic analyses also indicated strong activation of the KRAS-MAPK signaling pathway, suggesting that oncogenic signaling promotes the proliferation of gastric stem cells and drives the differentiation of intestinal-like epithelial lineages.
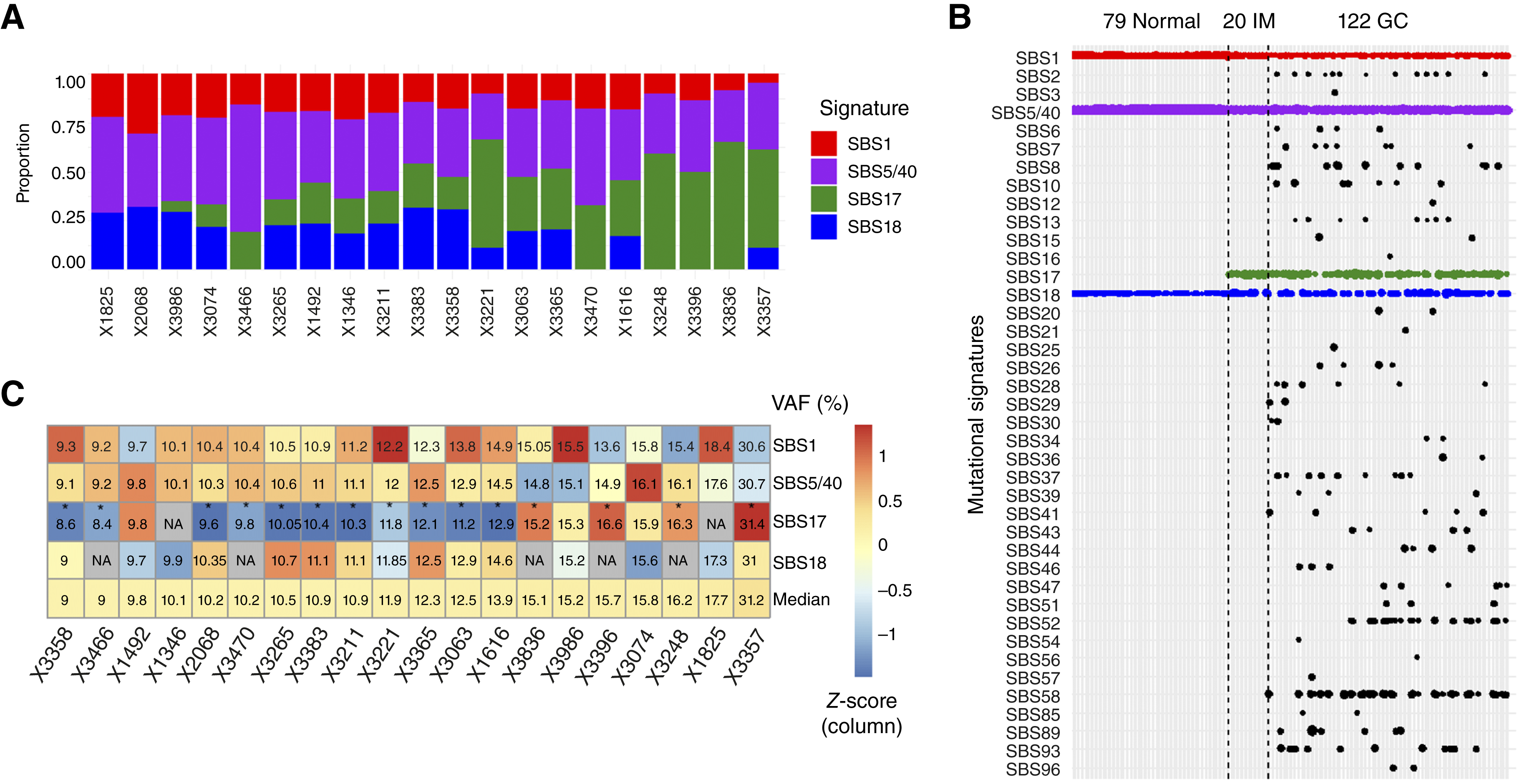
Figure 1. Mutational signatures in IM identified by whole-genome sequencing. (A) Mutational signatures in 20 IM samples detected by WGS, ordered by mutations attributed to SBS17. (B) Comparison across normal gastric tissue, IM, and gastric cancer, showing enrichment of SBS17 in IM and cancer. (C) Median variant allele frequencies (VAFs) of SBS mutations in IM samples; SBS17 mutations show lower VAFs, indicating early subclonal events.
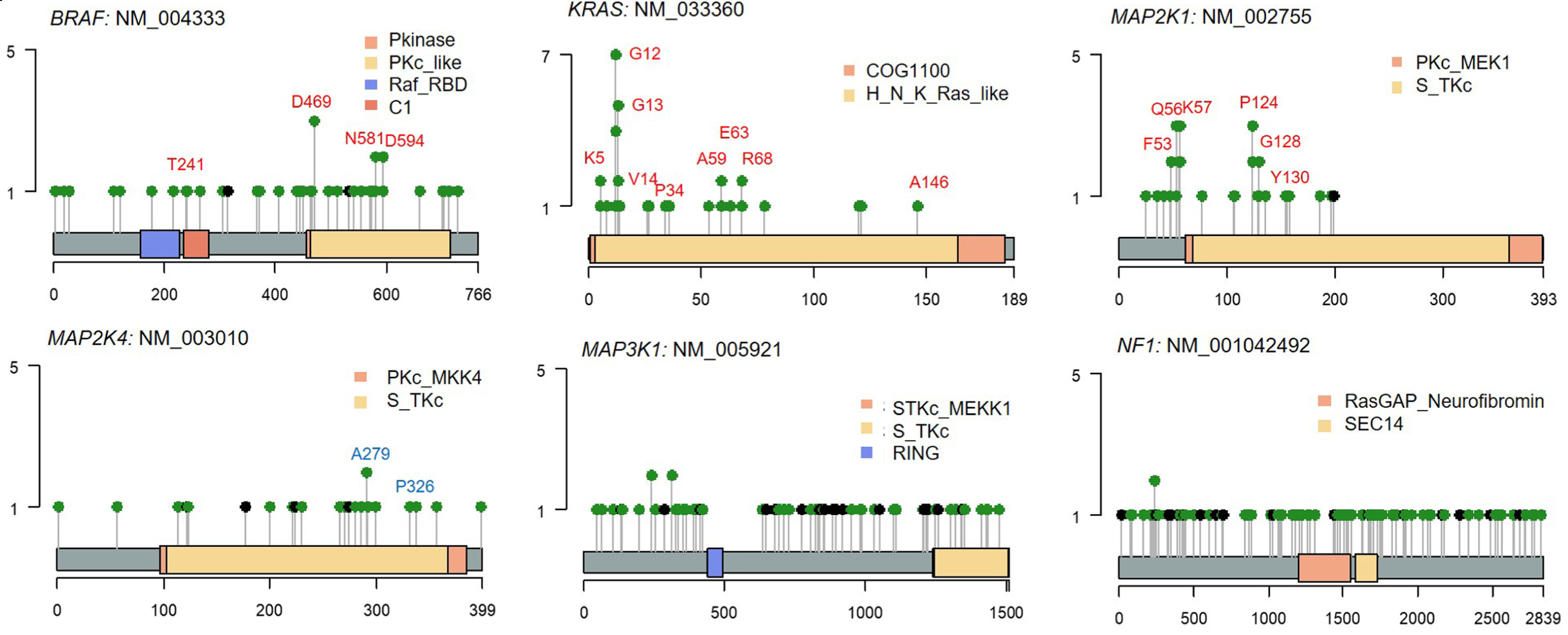
Figure 2. KRAS-MAPK pathway mutations in IM. Lollipop plots showing activating mutations in BRAF, KRAS, and MAP2K1 and inactivating mutations in MAP2K4, MAP3K1, and NF1 in KRAS-MAPK driver genes identified in IM. These alterations are associated with activation of KRAS-ERK signaling in IM.
To place these molecular alterations within their tissue context, the researchers further applied Stereo-seq OMNI Transcriptomics for FFPE Solution, enabling spatially resolved mapping of gene expression directly in clinical gastric cancer tissues. By capturing RNA molecules with spatial coordination barcoding (Coordinate ID, CID) on high-density patterned arrays, Stereo-seq allows transcriptomic signals to be reconstructed within intact tissue architecture, making it possible to investigate how genetic alterations and cellular states are organized across the gastric microenvironment.
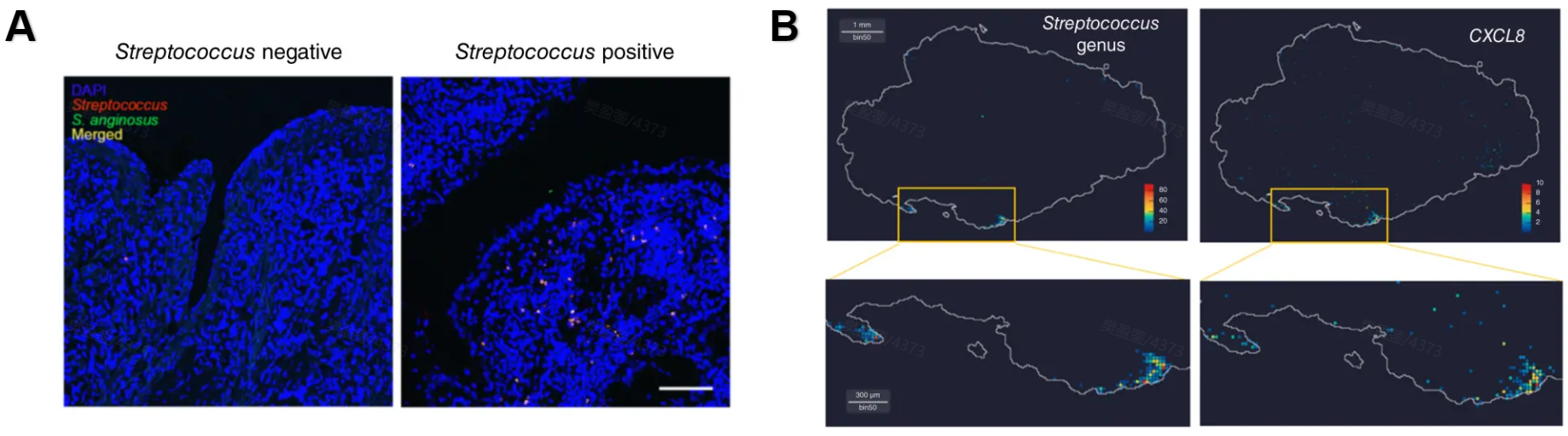
Figure 3. Spatial detection of Streptococcus in gastric cancer tissue. (A) FISH detection of Streptococcus (red) and Streptococcus anginosus (green) in gastric cancer tissue with DAPI nuclear staining (blue). (B) Spatial localization of Streptococcus co-localized with CXCL8 expression, revealed by STOmics Stereo-seq spatial transcriptomics (500 nm resolution)
STOmics' Stereo-seq Reveals Host-Microbe Interactions in Gastric Tissue
STOmics Stereo-seq proved critical for resolving the complex cellular architecture of the Correa cascade. With its 500 nm spatial resolution, Stereo-seq enables true cellular spatial transcriptomics, bridging the gap between genomic discoveries and their pathological context.
Conventional spatial platforms often struggle with archival FFPE tissues, where degraded RNA and low DV200 values limit transcript detection. Many rely on spot-based capture areas of 50-100 µm, averaging signals across heterogeneous cell populations. This "blurred lens" makes it difficult to determine whether inflammatory markers originate from metaplastic epithelial cells, immune infiltrates, or surrounding stroma. Importantly, such approaches often miss low-abundance microbial transcripts, leaving the host-microbe interface—a key driver of gastric disease progression—poorly resolved.
Using the STOmics Stereo-seq OMNI Transcriptomics for FFPE Solution, the study achieved spatial detection of both human and bacterial transcripts within the same tissue section. Researchers mapped the enrichment of Streptococcus anginosus in regions of inflammatory remodeling, linking microbial colonization with PIGR mutations, increased IgA⁺ plasma cells, and clonal hematopoiesis (CH)-associated immune dysregulation. These findings suggest that CH-driven immune imbalance may promote bacterial expansion and chronic inflammation, accelerating IM progression toward gastric cancer. Combined with H&E staining, Stereo-seq further resolved molecular gradients such as SOX9 expression, pinpointing the cellular niches where malignant transformation emerges—an evidentiary depth that conventional spatial technologies cannot achieve.
Figure 4. Spatial mapping of bacterial reads in gastric cancer tissue. Upper panel: Host cell-state architecture inferred from Stereo-seq data using the Stereopy square-bin clustering pipeline. Lower panels: Focal clusters of bacterial reads assigned to Streptococcus and Helicobacter. Color scales indicate bacterial read density. Stereo-seq's 500 nm resolution enables simultaneous spatial detection of host and microbial transcripts.
STOmics Matrix: Powering the Next Generation of Spatial Omics
Insights from studies like this spatial analysis of IM and gastric cancer progression highlight the growing importance of high-resolution spatial technologies. To support such research, STOmics provides a comprehensive portfolio of spatiotemporal omics solutions built on the Stereo-seq platform, enabling detailed exploration of tissue architecture and host-microbe interactions. Table 1 outlines the key technical specifications of these solutions along with their representative applications.
Table 1. STOmics product matrix
Future Horizons with Stereo-seq
As spatial biology reshapes cancer research, technologies like STOmics Stereo-seq can link genomic alterations to tissue architecture with nanometer resolution and clinical sample compatibility. In gastric cancer, spatial transcriptomics may identify early biomarkers, clarify host-microbe interactions, and reveal how driver mutations reshape local immune environments.
Bridging gastric carcinogenesis and precise clinical strategies, Stereo-seq enables PIs and lead researchers to uncover cellular interactions with unparalleled spatial resolution. Contact Us at info_global@stomics.tech for expert support in experimental design and advancing your research.
FAQs
Q: How does Stereo-seq handle low-quality FFPE samples?
A: Stereo-seq OMNI employs optimized hybridization and random priming strategies to capture fragmented RNA from archived FFPE samples, enabling robust gene detection even in specimens with low DV200 values.
Q: Why is 500nm resolution critical for microbiome studies?
A: Stereo-seq's ~500 nm DNA Nano Ball (DNB, a high-density DNA capture probe) resolution enables precise localization of individual transcripts, including low-abundance microbial signals. Unlike 50 µm spot-based methods that average signals, it resolves true cellular and subcellular origins of gene expression.
Q: Can I perform H&E and transcriptomics on the same slide?
A: Yes. Stereo-seq supports H&E staining and spatial transcriptomic profiling on the same tissue section, enabling direct registration of histomorphology with gene expression data.
Q: Why is spatial analysis important for gastric cancer research?
A: Cancer is a spatial battle. Stereo-seq provides the important comprehensive blueprint of the tumor microenvironment, revealing the hidden dependencies between mutated clones, immune cells, and microbes. Move beyond description into predictive insights for precision medicine.
