The human thymus functions as a biological "academy" where T cells develop and learn to distinguish self from non-self. Within this microenvironment, developing thymocytes interact with stromal cells—particularly thymic epithelial cells (TECs)—that guide positive and negative selection (Takahama, 2006). This process relies on a highly organized spatial architecture, especially in the thymic medulla, where rare epithelial populations mimic peripheral tissues to test developing T cells. Recent studies have identified mimetic TEC populations expressing lineage-specific transcription factors, highlighting the importance of understanding their spatial organization within the thymus (Anderson & Su, 2016).
Limitations of Existing Spatial Approaches
Traditional spatial technologies often force researchers into a compromise: either accept low resolution that averages multiple cells together, or settle for a tiny field of view that misses organ-scale structural organization (Marx, 2021).
These limitations present several key challenges for thymic research. First, rare cell populations such as mimetic TECs, which represent less than 0.4% of thymic cells, are extremely difficult to detect without high-resolution spatial profiling. Second, the loss of spatial context means interactions between cortical and medullary microenvironments cannot be fully resolved (Longo, 2021). Finally, the limited field of view of many spatial platforms prevents the preservation of broader anatomical context in large organs like the thymus. A new generation of spatial technologies capable of combining nanoscale resolution with centimeter-scale tissue coverage is therefore essential for advancing our understanding of thymic architecture.
Spatial Profiling Strategy with STOmics' Stereo-seq LCD and Stereo-CITE Solution
To construct a comprehensive spatial atlas of the human thymus, the researchers designed a multi-modal workflow integrating several genomic technologies, as reported by Kamaraj et al. in Nature Communications on March 10, 2026 (Kamaraj et al., 2026). Human fetal and pediatric thymus samples were cryosectioned and processed using Stereo-seq spatial transcriptomics, which captures mRNA directly from tissue sections through DNB (DNA Nano Ball) with spatial coordination barcoding (Coordinate ID, CID). This approach enabled high-resolution transcriptomic mapping while preserving native tissue architecture.
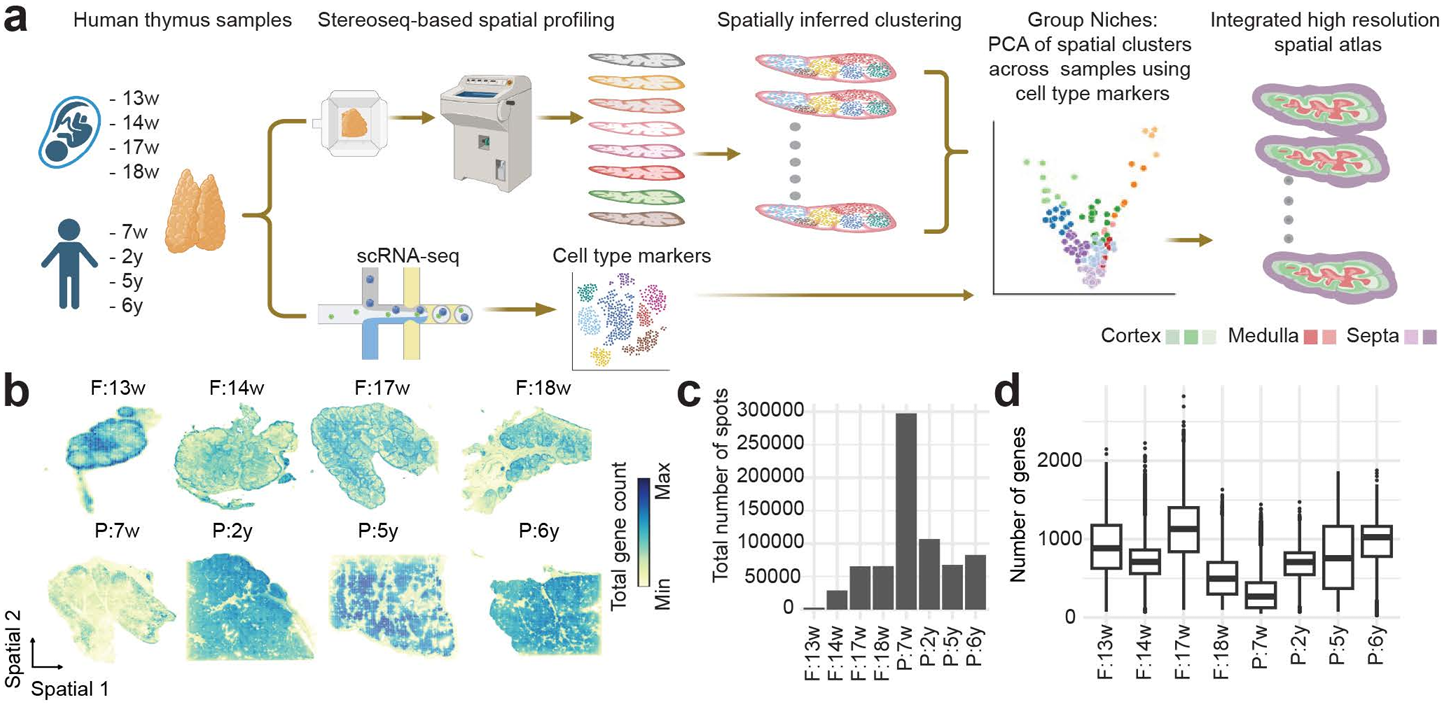
Figure 1. STOmics' Stereo-seq and scRNA-seq powered high-resolution spatial atlas of the human thymus, mapping cortical-medullary architecture and diverse immune niches from 13 weeks post-conception through childhood.
A key advantage of the study was the use of Stereo-seq Large Chip Designs (LCD), including formats up to 2 × 3 cm. In developmental biology, spatial context is essential; analyzing small tissue fragments can obscure broader architectural transitions such as the cortico-medullary junction. The large capture area allowed researchers to map extensive regions of fetal and pediatric thymus and observe how different mimicry programs—neural, muscular, and pancreatic lineages—are spatially organized within the medulla.
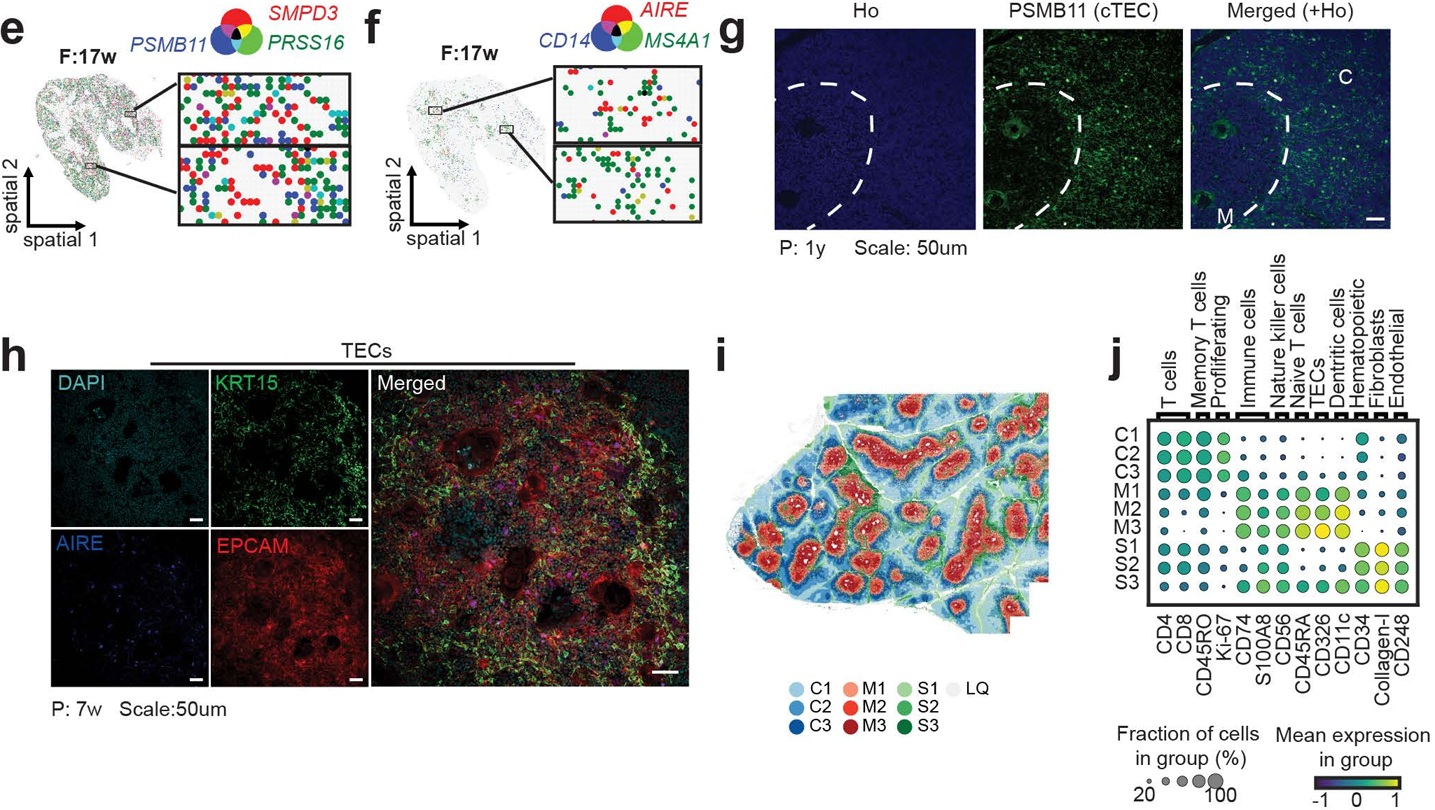
Figure 2. Spatial geopositioning of human thymic niches across age. STOmics' Stereo-seq resolves anatomical regions via marker expression: PRSS16+ cortex (cTECs/DP T cells), AIRE+ medulla (mTECs/DCs/B cells), and COL1A2/VWF/CD34+ septa, revealing functional segregation at high resolution.
The spatial dataset was further integrated with single-cell RNA sequencing for cell type annotation and multiplexed protein imaging using MACSima. To simultaneously detect RNA and protein markers in the same tissue section, the team applied Stereo-CITE proteo-transcriptomics. This dual-omics strategy confirmed that transcription factors such as INSM1, ASCL1, and MYOG mark coordinated mimetic programs that expose developing T cells to a molecular representation of peripheral tissues.
The Utility and Significance of Stereo-seq
Stereo-seq is critical to this study because it addresses fundamental limitations that have long constrained thymus research. Traditional single-cell approaches, while informative, disrupt tissue architecture and cannot resolve the spatial relationships between rare cell populations and their microenvironment (Ståhl et al., 2016).
Stereo-seq leverages two key capabilities of the STOmics platform. First, its ultra-high spatial resolution (down to 500 nm) enables detection of transcriptional signals from extremely small cellular niches, making it possible to identify rare mimetic TECs expressing transcription factors such as AIRE, FEZF2, ASCL1, and INSM1—populations that occur at frequencies below 0.4% and are easily missed by lower-resolution platforms.
Figure 3. Medullary mimeTF mapping. STOmics' Stereo-seq resolves rare mimetic TECs and TFs (ASCL1, MYOG) within EPCAM+ niches. Multi-omic validation confirms this spatial logic, decoding the mechanisms of thymic mimicry and central tolerance.
Second, the integration of Stereo-CITE multi-omics profiling allows simultaneous detection of RNA and protein markers within the same tissue section, providing multi-layer validation that confirmed mimetic TECs cluster within specialized KRT1-high regions near Hassall’s corpuscles, forming a spatial hub for antigen presentation and immune tolerance. As a result, the study uncovered 70 transcription factors associated with mimetic TEC regulation, revealing previously unknown regulatory networks within the thymic microenvironment.
Figure 4. Functional diversity and spatial enrichment of mimeTFs. Rare mTECs expressing lineage-specific TFs reside in KRT1+ clusters and show significant spatial proximity to B cells and dendritic cells, facilitating the presentation of non-thymic self-antigens within the medullary microenvironment.
Beyond these technical advances, Stereo-seq enables a paradigm shift from descriptive biology to functional spatial genomics. The ability to map large tissue sections across developmental stages—from 13 weeks post-conception to early childhood—provides a foundation for longitudinal studies of thymic maturation and involution.
STOmics' Matrix for High-Resolution Mapping
Studies such as this spatial mapping of the human thymus highlight the need for technologies capable of resolving rare cell populations within complex tissue architectures. To support such research, STOmics provides a comprehensive portfolio of spatiotemporal omics solutions based on the Stereo-seq platform, enabling molecular mapping at nanometer-scale resolution across centimeter-scale tissue regions. Table 1 summarizes the key technical specifications of these solutions and their representative applications in spatial biology.
Table 1. STOmics product matrix
The Future of Spatial Biology
As precision medicine increasingly demands cellular resolution of tissue microenvironments, STOmics' Stereo-seq establishes a new standard for understanding how spatial organization governs immune function, autoimmune risk, and age-related thymic decline. Looking forward, the platform’s robust performance supports the transition from 2D mapping to 3D organ-scale reconstruction, offering researchers a powerful tool to investigate fundamental questions in immunology with unprecedented spatial resolution. To explore how Stereo-seq can advance your spatial biology research, please feel free to Contact Us or email info_global@stomics.tech.
FAQs
Q: Why is 500 nm resolution considered the suitable standard for rare cell research?
A: Most spatial technologies use "bins" or "spots" that are 10-50 μm in diameter, which typically capture 10 to 30 cells at once. At 500 nm, Stereo-seq captures data at a cellular level, ensuring that the unique signal of a rare cell (like an mTEC) isn’t drowned out by its more common neighbors.
Q: How does the Stereo-seq LCD impact batch effects and data consistency?
A: By using chips as large as 13 × 13 cm, researchers can place multiple samples or entire organ sections on a single chip. This ensures identical processing conditions across all samples, virtually eliminating the experimental noise and batch effects commonly found when stitching together many smaller images.
Q: Can I integrate my existing scRNA-seq data with Stereo-seq results?
A: Yes. Stereo-seq data is highly compatible with standard bioinformatics pipelines. Most researchers use their scRNA-seq data as a "reference atlas" to perform cell-type annotation transfer, allowing them to map their identified clusters back onto the high-resolution spatial coordinates provided by Stereo-seq.
References
