Soybeans are one of the world's most important crops, providing the majority of plant protein and oil for human consumption and livestock feed. Facing ever-increasing global demand, soybean production is projected to double by 2050 compared to 2015 levels (Hartman et al., 2011). To support this growth, developing more resilient and higher-yielding varieties has become a central goal in crop science.
Achieving this requires a deeper understanding of the genetic networks that regulate organ development. As a typical legume, soybean forms specialized root nodules that enable nitrogen fixation—key biological processes for yield improvement (Roy et al., 2019). However, the precise regulatory mechanisms underlying these complex organs remain largely unknown. Capturing their spatial and cellular complexity requires not only traditional transcriptomics but also approaches that preserve spatial context.
The Persistent Gaps in Plant Transcriptomics
Despite advances in bulk RNA-seq and snRNA-seq, key limitations remain. Bulk RNA-seq averages signals across tissues, masking cellular heterogeneity. While snRNA-seq can distinguish cell types, it inevitably loses spatial information about cells within tissues (Longo et al., 2021). These limitations are particularly pronounced in plants, where rigid cell walls and complex tissue organization complicate cell isolation. As a result, critical biological questions—such as how specific genes coordinate spatially during nodule formation or leaf development—have remained unresolved.
Building a Multi-Omics Atlas: Integrating Spatial Context
To overcome these limitations and dissect gene regulation during soybean organ development, a study published in Molecular Plant in 2025 constructed a comprehensive integrated transcriptome map in the soybean field to date (Fan et al., 2025). It established an integrated framework combining bulk RNA-seq, snRNA-seq, and spatial transcriptomics viaStereo-seq. Bulk RNA-seq across 314 samples provided a global expression landscape, while snRNA-seq resolved cellular heterogeneity and identified distinct cell clusters.
Stereo-seq was then applied to cryosectioned tissues including root tips, nodules, leaves, stems, and shoot apices, enabling spatially resolved transcriptome profiling. Each section captured 13,733-37,086 genes, with high gene counts per spatial bin, ensuring both sensitivity and spatial precision. By integrating snRNA-seq with STOmics' Stereo-seq Transcriptomics Solution, the study reconstructed spatial single-cell atlases, mapping cell types back to their anatomical locations.
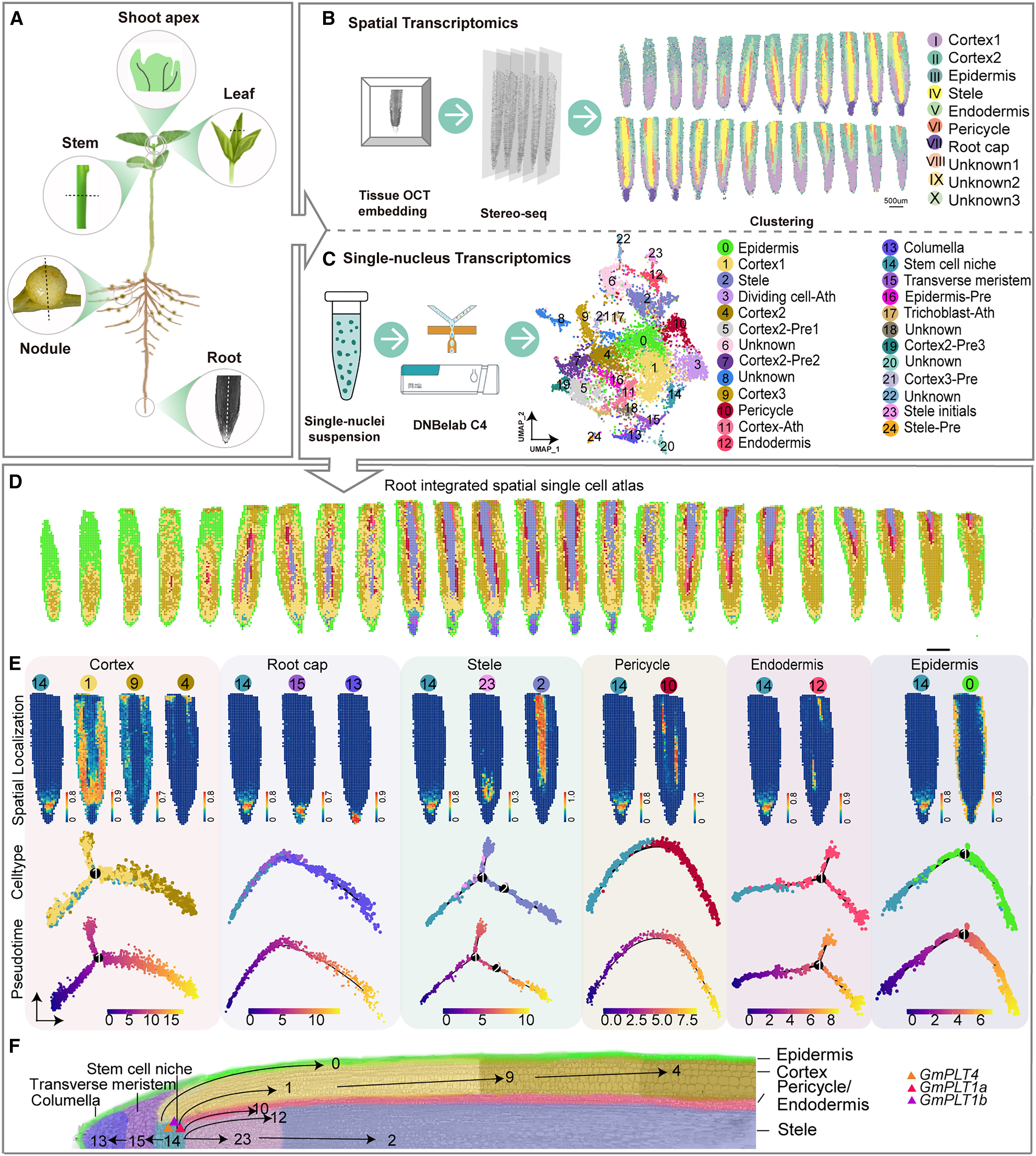
Figure 1. Spatial single-cell atlas of soybean roots. (A) Overview of organs analyzed by Stereo-seq and snRNA-seq. (B) Stereo-seq reveals 10 spatial clusters in roots. (C) snRNA-seq identifies 25 root cell clusters. (D) Integration maps cell types to spatial locations. (E) Pseudotime analysis shows root cell differentiation trajectories. (F) Model of root cell development highlighting GmPLT expression.
This approach enabled developmental trajectory analysis, showing how stem cell niches differentiate into multiple root tissues, regulated by transcription factors such as GmPLT1a, GmPLT1b, and GmPLT4. In nodules, spatial clustering revealed distinct vascular regions and regulatory programs. Key genes including GmHB1a, GmHB1b, and GmHB21 were precisely localized, and functional validation confirmed their roles in nodule formation. Additionally, GmPMT1-GmPMT6 were identified as nodule-specific transporters controlling sugar allocation during symbiosis. These findings demonstrate how spatial transcriptomics enables the interpretation of gene expression in a mechanistic context.
Figure 2. Spatial localization of key genes and their role in nodule development. (A) Cell type-specific expression of functionally validated SNF genes, mapped using Stereo-seq. (B) Phylogenetic tree of GmHB1a, GmHB1b, and GmHB21. (C) Localization of GmHB1a and GmHB21 in nodule tissues, with distinct expression in vascular junction and nodule vascular cells.
How Stereo-seq Solves Critical Limitations in Plant Spatial Biology
In this study, Stereo-seq demonstrated its core advantages in revealing the heterogeneity of soybean organ development. Without spatial transcriptomics, it is difficult to define the precise spatial boundaries of specific genes—such as the sugar transporter family GmPMTs. Using Stereo-seq, the study accurately identified that GmPMT3 and GmPMT5 are primarily expressed in the symbiotic zone of root nodules, indicating their key roles in coordinating carbon allocation to microbes. Furthermore, in the root stem cell niche, Stereo-seq successfully mapped the spatial patterns of the GmPLT gene family within the quiescent center and surrounding initial cells. This spatial mapping capability not only improves the accuracy of cell-type annotation but also reveals developmental trajectories and spatial associations that bulk or dissociated single-cell methods cannot capture.
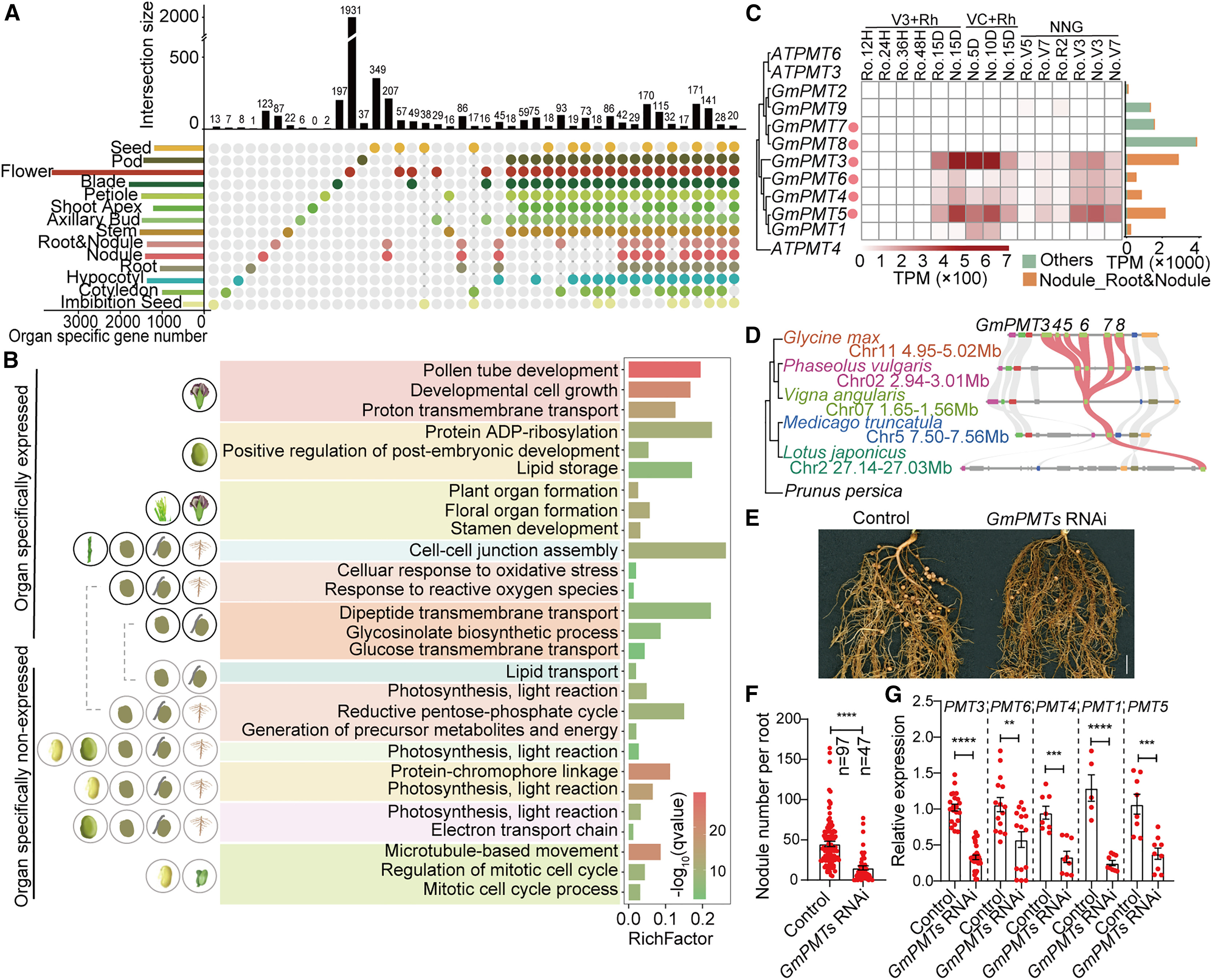
Figure 3. Nodule-specific expression and functional analysis of GmPMT genes in soybean. (A) Upset plot showing organ-specific gene intersections. (B) GO terms enriched for genes specifically expressed in nodules, including glucose transport. (C) Heatmap of GmPMT expression in nodules and roots. (D) Synteny analysis of GmPMT genes across legume species. (E-G) RNAi knockdown of GmPMTs reduced nodule number (E, F) and gene expression (G), confirming their role in nodule development.
By linking gene expression with spatial organization and function, the STOmics' Stereo-seq Transcriptomics Solution not only overcomes the difficulties faced by traditional techniques but also provides a powerful solution for decoding complex plant systems, enabling whole-transcriptome mapping within complete tissues. With a resolution of 500 nm and a field of view down to the centimeter level, it uniquely combines nanoscale precision with full organ coverage. Its compatibility with multiple sample types and scalable design further make it a key technology for advancing plant functional genomics and crop research.
STOmics Matrix: Enabling High-Resolution Spatial Transcriptomics
Studies such as this spatial transcriptomic atlas of soybean organ development highlight the need for technologies capable of resolving cell-type heterogeneity and spatial gene regulation within complex plant tissues. Beyond soybean nodule analysis, STOmics offers an adaptable portfolio covering diverse plant biology and functional genomics needs. Table 1 summarizes the key technical specifications of these solutions and their representative applications in plant spatial biology and crop research.
Table 1. STOmics product matrix
Looking Ahead: Scaling Spatial Omics in Plant Science
STOmics' Stereo-seq is redefining plant functional genomics by enabling high-resolution, spatially resolved whole-transcriptome mapping. It empowers researchers to uncover key regulatory pathways and link gene activity to precise developmental contexts, supporting more targeted crop breeding strategies. As spatial omics advances, plant research is entering a new data-driven era.
If your team is working on plant development or has similar spatial biology needs, STOmics would be happy to support your research. To explore how Stereo-seq can advance your work, please feel free to Contact Us or email info_global@stomics.tech.
FAQs
Q: What is the specific significance of Stereo-seq's 500 nm resolution for plant tissue research?
A: While most spatial technologies operate at the micron level—where a single sampling spot often encompasses multiple cells or cellular fragments—Stereo-seq's 500 nm resolution enables capture at a sub-cellular scale. This allows for the precise resolution of intricate and densely packed cellular architectures, such as the stem cell niches in soybean root tips, improving the accuracy of mapping transcriptomic signals to their original spatial locations.
Q: Is Stereo-seq robust enough for challenging or degraded plant tissues?
A: Yes. The workflow supports both Fresh Frozen (FF) sections and non-model species. For archival or low-quality materials (low RIN/DV200), the Stereo-seq OMNI solution maintains high sensitivity at 500 nm resolution, allowing researchers to retrospectively analyze complex developmental or pathological mechanisms in tissues previously considered inaccessible.
Q: How does spatial transcriptomics accelerate precision crop improvement?
A: By mapping the whole transcriptome across cm-scale fields of view, Stereo-seq identifies the precise spatial-temporal activity of regulatory genes. This provides actionable, cell-type-specific targets for CRISPR/Cas9 editing and marker-assisted selection—transforming broad genomic observations into a "molecular design" approach for high-yield, resilient varieties.
References
- Fan, J. et al. A large-scale integrated transcriptomic atlas for soybean organ development. Molecular Plant (2025).
- Hartman, G. L., West, E. D. & Herman, T. K. Crops that feed the World 2. Soybean—worldwide production, use, and constraints caused by pathogens and pests. Food Security 3, 5–17 (2011).
- Roy, S. et al. Celebrating 20 Years of Genetic Discoveries in Legume Nodulation and Symbiotic Nitrogen Fixation. The Plant Cell 32, 15–41 (2019).
- Longo, S. K., Guo, M. G., Ji, A. L. & Khavari, P. A. Integrating single-cell and spatial transcriptomics to elucidate intercellular tissue dynamics. Nature Reviews Genetics 22, 627–644 (2021).
