Liver metastasis (LM), particularly from colorectal cancer, remains a major challenge. Beyond direct cytotoxicity, chemotherapy reshapes the liver's immune architecture (Galluzzi et al., 2016). Within this immune-tolerant organ, resident Kupffer cells (KCs) exhibit high plasticity and can be reprogrammed by chemotherapy (Jenne & Kubes, 2013). Emerging evidence suggests that instead of acting as sentinels of anti-tumor immunity, these KCs are subverted into barriers that suppress immune responses. Understanding how chemotherapy shifts KCs from guardians to enablers of immune evasion is essential for improving therapeutic strategies in colorectal liver metastases (CRLMs).
Current Limitations in Spatial and Single-cell Research
Investigations into the tumor microenvironment are hindered by a critical technical bottleneck: the inability to balance high resolution with a comprehensive field of view. Conventional approaches lack sufficient resolution: scRNA-seq loses spatial context, while early spatial methods aggregate multiple cells, obscuring precise immune–tumor interactions (Ståhl et al., 2016; Liu et al., 2020).
These limitations become particularly critical when studying rare or spatially restricted populations, such as LEPR⁺ KCs, whose functional roles depend on precise positioning relative to tumor cells, dying cells, and infiltrating T cells; without sufficient spatial resolution, these closely interacting cell populations are easily averaged together, obscuring key immune–tumor interactions and masking localized immunosuppressive niches. Researchers have thus struggled to pinpoint exactly where these cells reside in relation to chemotherapy-induced immunogenic cell death (ICD) signals, making it difficult to capture their dynamic redistribution across whole-lesion architecture and tumor margins simultaneously.
Integrating Multi-Omics for Spatiotemporal Insights with Stereo-seq
To bridge the gap between cellular identity and spatial function, a groundbreaking study by Wang et al. (2026), recently published in Cancer Cell, employed a sophisticated multi-omic strategy, integrating single-cell RNA sequencing (scRNA-seq) with the STOmics' Stereo-seq Transcriptomics Solution. The study utilized a diverse array of models, including human CRLM clinical samples and an AKP-derived murine liver metastatic model. By treating these models with standard chemotherapy, the researchers created a timeline of tissue remodeling to track the lineage plasticity of KCs from a steady state to a post-treatment pathological state.
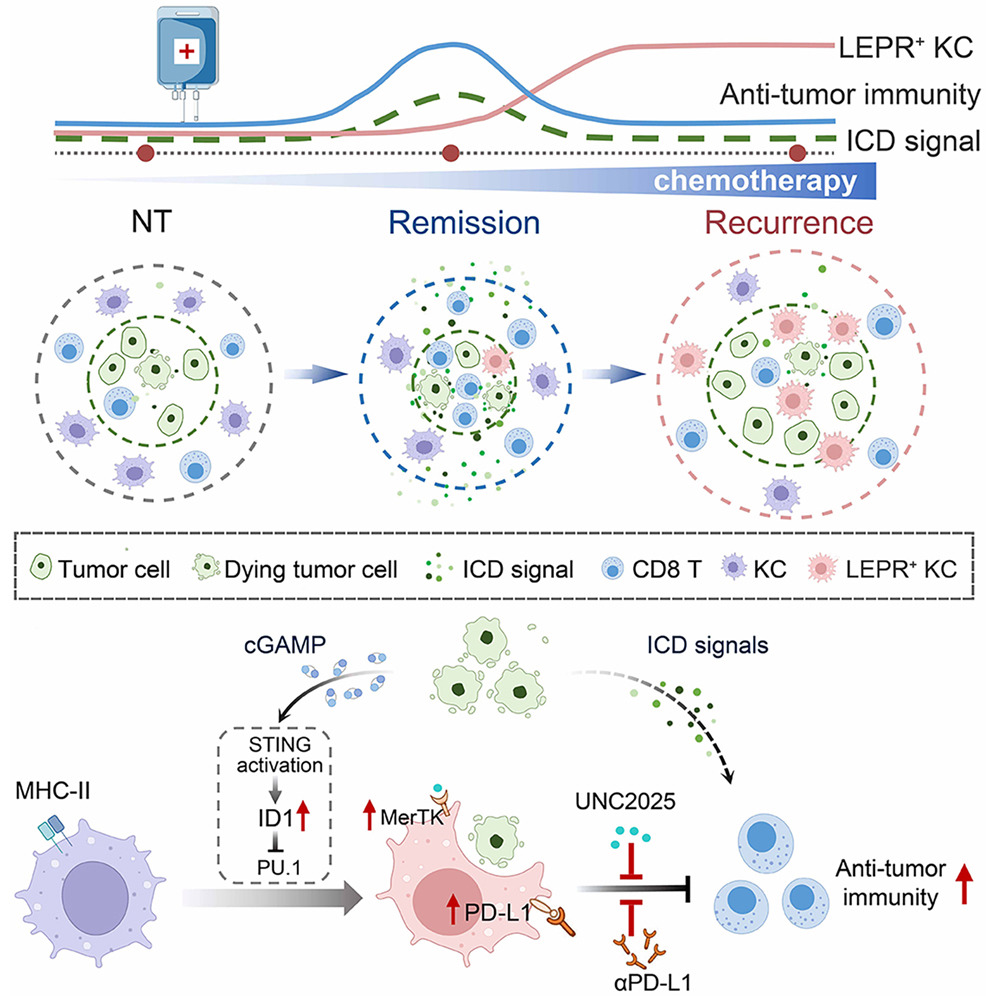
Figure 1. Chemotherapy not only activates antitumor immunity but also reprograms Kupffer cells (KCs) toward an immunosuppressive LEPR+ state via STING signaling driven by tumor-derived cGAMP. These KCs infiltrate tumors and promote immune evasion and chemoresistance through MerTK-dependent efferocytosis. Targeting LEPR+ KCs restores tumor immunogenicity and enhances T cell responses.
The workflow centered on the ability of Stereo-seq to provide 500nm resolution across a centimeter-scale field of view. This allowed the team to perform "neighborhood analysis" at a sub-cellular level. By precisely mapping the spatial coordinates of transcripts, they successfully correlated the paracrine release of cGAMP from dying tumor cells—a byproduct of chemotherapy-induced DNA damage—directly with the activation of the STING signaling in nearby resident KCs.
The nanometer-scale precision of Stereo-seq enabled the researchers to observe the step-by-step differentiation process. They identified that chemotherapy-driven signals trigger the recruitment of the transcription factor PU.1, which binds to the LEPR promoter, effectively transforming "bystander" KCs into a highly specialized, pro-tumorigenic LEPR+ subset. This meticulous mapping proved that these cells do not just exist within the liver; they specifically relocate and cluster within the tumor parenchyma following treatment. By capturing the entire section of the liver metastases, the large-scale capture area ensured that these spatial correlations were statistically significant across the whole organ, providing a truly global perspective on immune evasion.
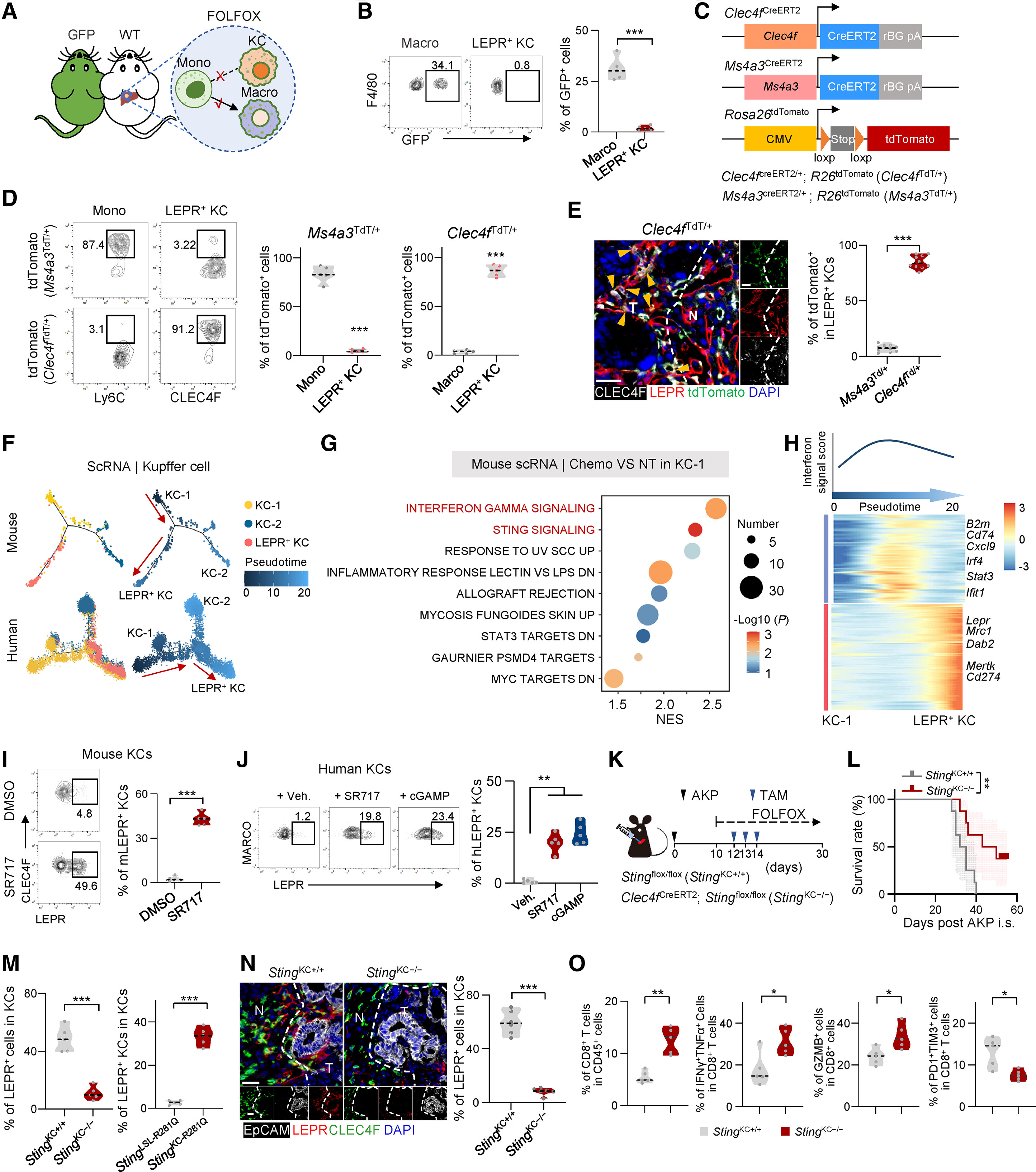
Figure 2. Chemotherapy-induced STING signaling drives LEPR+ KC differentiation. Lineage tracing confirmed that LEPR+ KCs originate from resident liver KCs via tumor-derived cGAMP and STING signaling. In models, blocking this pathway inhibited LEPR+ KC formation, restored CD8+ T cell infiltration, and extended survival, identifying a key spatial target to overcome chemoresistance.
Why Spatial Resolution Matters: Stereo-seq in Action
In this study, the STOmics' Stereo-seq Transcriptomics Solution is integrated into the workflow to map KC dynamics within tumor tissues, enabling spatial comparison between chemo-sensitive and resistant regions. This allows researchers to directly investigate how LEPR⁺ KCs redistribute and interact with tumor and immune cells in situ.
Without high-resolution spatial tools, it would be difficult to determine whether LEPR⁺ KCs infiltrate tumors or remain peripheral, or to link their distribution with CD8⁺ T cell exclusion and chemotactic signals such as ATP-P2RY2. By providing 500 nm resolution, Stereo-seq reveals that LEPR⁺ KCs accumulate within tumor regions during relapse and are spatially associated with localized immune suppression. At the same time, its centimeter-scale field of view enables whole-lesion analysis in a single section, capturing tumor core, margins, and microenvironment while reducing batch effects. By localizing key genes such as MERTK, P2RY2, and ID1, Stereo-seq connects transcriptional programs with spatial function, offering a clearer mechanistic understanding of how chemotherapy-driven KC reprogramming promotes immune evasion.
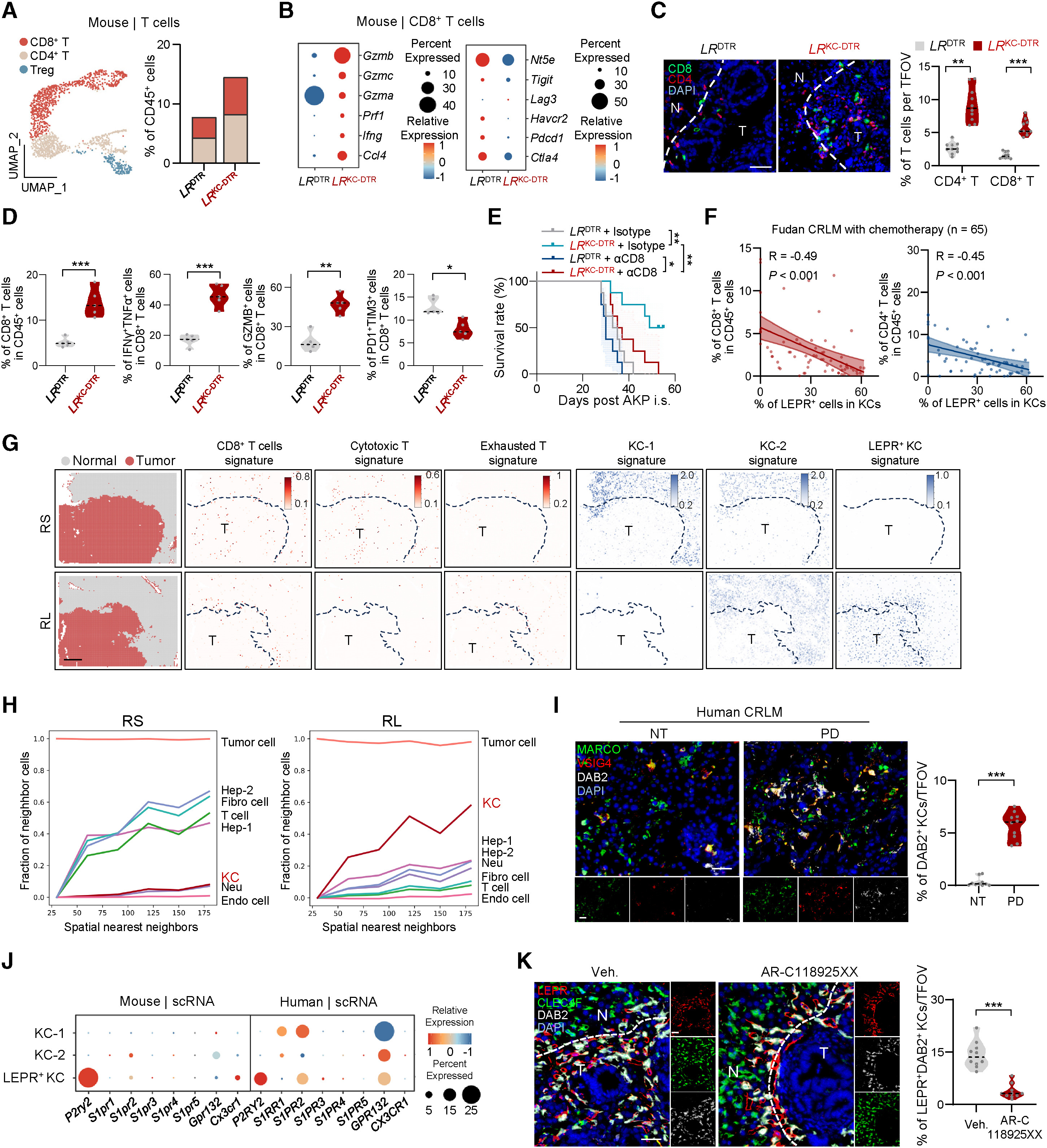
Figure 3. LEPR+ KCs infiltrate tumors and suppress antitumor immunity. Integrating T-cell UMAPs and clinical correlations, researchers used Stereo-seq to map LEPR+ KCs within the tumor parenchyma. Neighborhood analysis confirmed that these cells—marking high expression of DAB2 and MerTK—physically infiltrate the tumor during chemoresistance, negatively correlating with CD8+ T cell presence and overall survival.
Empowering Oncology Research: The STOmics Spatial Portfolio
As demonstrated by this groundbreaking spatial transcriptomic atlas of liver metastases, modern oncology demands technologies that can bridge the gap between complex immune plasticity and the physical shielding mechanisms within the tumor microenvironment. To empower researchers in deciphering these intricate cellular dialogues, we offer a versatile portfolio of spatiotemporal omics solutions powered by the Stereo-seq platform.
Table 1. STOmics product matrix
The Future Outlook from Discovery to Clinical Intervention
The success of this study underscores the future of precision medicine. By defining the spatial coordinates of resistance markers like LEPR, STING1, and MerTK, Stereo-seq provides a roadmap for "spatial-targeted" therapies. If your laboratory is investigating tumor-immune dynamics, liver pathology, or the spatial mechanisms of drug resistance, Stereo-seq offers the resolution and scale necessary to uncover the next generation of therapeutic targets. To explore how our spatiotemporal solutions can accelerate your specific research in oncology or clinical immunology, please feel free to Contact Us or reach out via email at info_global@stomics.tech.
FAQs
Q: Why is spatial transcriptomics essential for studying tumor immunity?
A: Tumor immunity depends on cell location and interaction, not just gene expression. Traditional methods lose spatial context, making it hard to resolve processes like immune infiltration or exclusion. Spatial transcriptomics preserves tissue architecture, enabling direct mapping of immune–tumor interactions in situ.
Q: What makes Stereo-seq different from other spatial platforms?
A: Stereo-seq combines 500 nm resolution with centimeter-scale coverage, delivering both near-subcellular precision and whole-tissue analysis in one experiment. Its high sensitivity and large capture area allow more accurate detection of cell interactions while reducing batch effects.
Q: What insights did spatial analysis uniquely reveal in this study?
A: It showed that LEPR+ Kupffer cells infiltrate tumor regions during chemoresistance and are spatially linked to reduced T cell activity. This directly connects their location with immune suppression, revealing mechanisms that non-spatial methods cannot capture.
