Cervical cancer remains a leading cause of malignancy-related mortality among women globally (Sung et al., 2021). While primary treatments often involve microtubule-targeting agents like paclitaxel, the emergence of chemotherapy resistance poses a formidable challenge to clinical success. Recent breakthroughs have shifted focus toward the "epitranscriptome"—the collection of chemical modifications on RNA molecules—specifically RNA 5-methylcytosine (m5C) modifications. These post-transcriptional markers are now recognized as pivotal regulators of gene expression, influencing everything from mRNA stability to protein translation (Song et al., 2022). Understanding how these modifications are distributed within the tumor microenvironment is essential for identifying the next generation of therapeutic targets and overcoming drug evasion.
The Spatial Blind Spot in Modern Oncology
Despite known drug resistance mechanisms like efflux pump upregulation, many chemotherapy-resistant cases remain unexplained because of the inability to study gene regulation in its spatial context. A key limitation is the inability to map post-transcriptional regulatory events while preserving tumor spatial architecture. Bulk RNA sequencing masks intratumoral heterogeneity, while scRNA-seq destroys spatial context altogether (Ståhl et al., 2016). Researchers need tools that capture high-resolution transcriptomic data with intact morphology to pinpoint where resistance pathways activate. This is critical for spatially restricted phenomena—such as NSUN2⁺, YBX1⁺, or SERPINB5⁺ cell enrichment at tumor boundaries, as demonstrated later in this study—and for profiling entire tumor sections to reveal spatial gradients and microenvironmental interactions. Overcoming this spatial blind spot is essential to understanding chemoresistance and cancer progression.
Resolving Malignant Subpopulations through Multi-Omics Spatial Mapping with Stereo-seq
To systematically dissect m5C-driven gene regulation, Liu et al. (2026) applied four complementary sequencing strategies to compare tumor and normal tissues from multiple molecular dimensions. RNA BS-seq was used to characterize m5C methylation differences, revealing widespread hypermethylation in cancer, with m5C sites preferentially enriched within coding sequences. In parallel, RNA-seq profiled transcriptional differences, identifying key upregulated genes such as SERPINB5⁺, TP63⁺, and HK2⁺. scRNA-seq further resolved cellular heterogeneity, distinguishing malignant and non-malignant cell populations, while spatial transcriptomics captured spatial distribution differences, preserving tissue architecture and mapping gene expression back to its in situ location. Through this integrated design, the study linked epitranscriptomic modification, gene expression, and spatial organization, ultimately revealing coordinated regulatory changes underlying tumor progression.
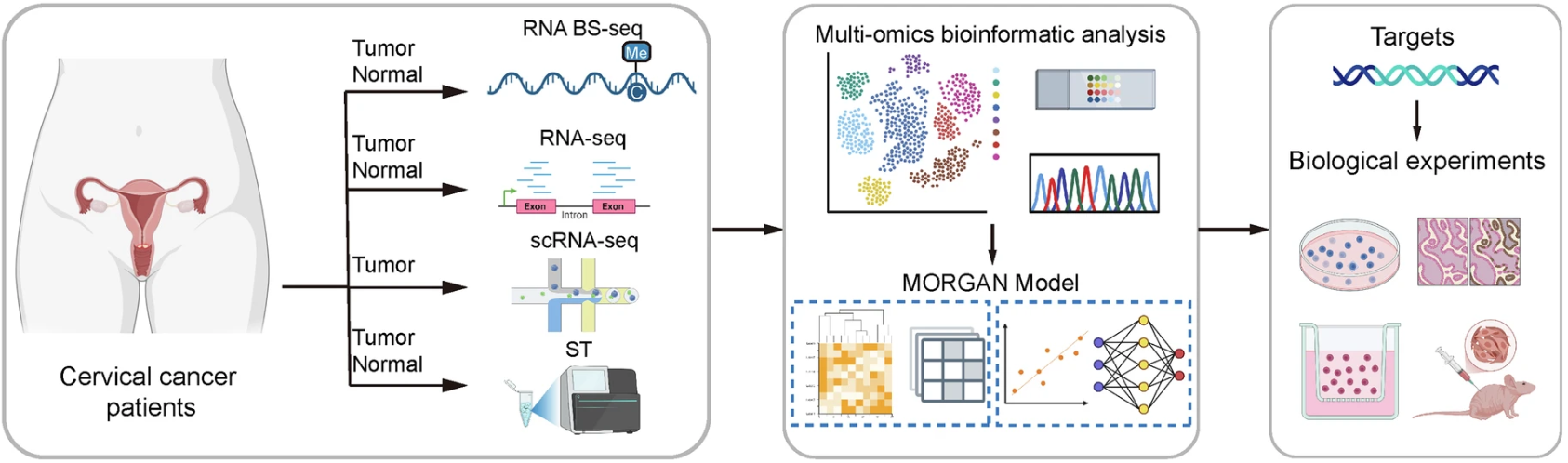
Figure 1. Schematic of the integrated spatial-epitranscriptomic workflow. Tumor and normal cervical tissues were profiled using RNA BS-seq and bulk RNA-seq, with tumor samples further analyzed via scRNA-seq and high-definition stereo-seq. Data integration through the MORGAN model facilitated the identification of oncogenic targets, followed by rigorous experimental validation.
Importantly, the introduction of spatial omics enabled the identification of a core regulatory axis centered on NSUN2(writer), YBX1 (reader), and SERPINB5 (effector), along with downstream cell cycle–related genes such as CENPE, CDK1, and CCNB1. Spatial mapping showed that NSUN2⁺ and YBX1⁺ cells are enriched at the tumor periphery, while SERPINB5⁺ cells form a distinct malignant subpopulation with enhanced proliferative capacity. Subsequent functional validation—including gene knockdown and RNA stability assays—confirmed that m5C modification mediated by NSUN2stabilizes SERPINB5 mRNA via YBX1, thereby promoting tumor growth and conferring resistance to chemotherapeutic agents such as paclitaxel and vincristine. Together, these findings demonstrate how integrating multi-omics with spatial resolution enables a more complete understanding of epitranscriptomic regulation and its functional impact in cancer.
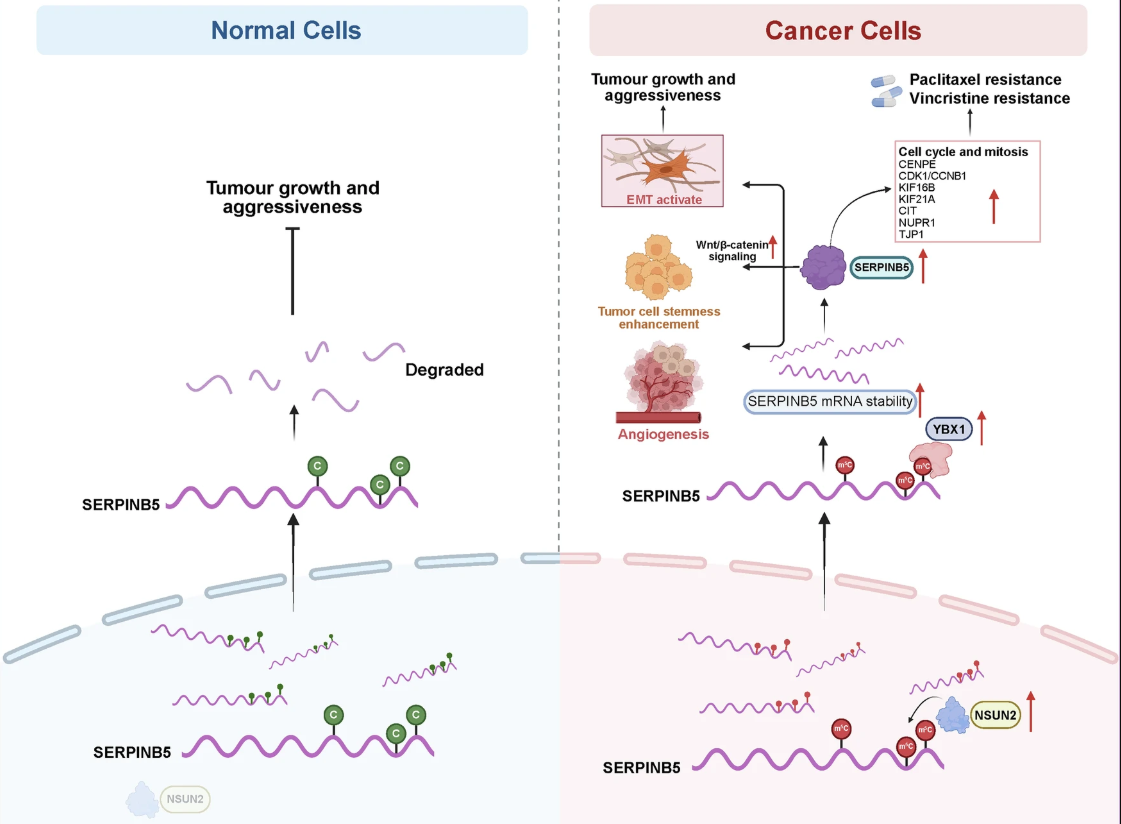
Figure 2. Mechanistic model of the NSUN2-SERPINB5 axis. In cancer cells, NSUN2-mediated m5C methylation and YBX1 binding stabilize SERPINB5 mRNA. This drives stemness, angiogenesis, and EMT, while upregulating mitotic and cytoskeletal regulators (CDK1, CCNB1, KIF16B) to fuel proliferation and evade chemotherapy-induced arrest.
Stereo-seq Advantage: From Spatial Resolution to Functional Insight
In this study, the introduction of the Stereo-seq transcriptomics solution was critical for translating molecular findings into spatially meaningful biological insights. From the perspective of lesion-associated gene expression, Stereo-seq enabled panoramic scanning of entire tumor sections with 500 nm resolution and a centimeter-scale field of view. This allowed researchers to precisely localize abnormal expression patterns within the tumor architecture.
The data revealed that NSUN2⁺ and YBX1⁺ cells are highly enriched at the invasive tumor periphery, highlighting a spatially restricted activation of the m5C regulatory machinery specifically at disease-relevant regions. In contrast, SERPINB5 expression was confined to defined malignant subpopulations within cancer-cell-rich areas. Such findings demonstrate that gene dysregulation is not uniform across the tumor but closely linked to specific pathological niches—insights that would be obscured without high-resolution spatial mapping.
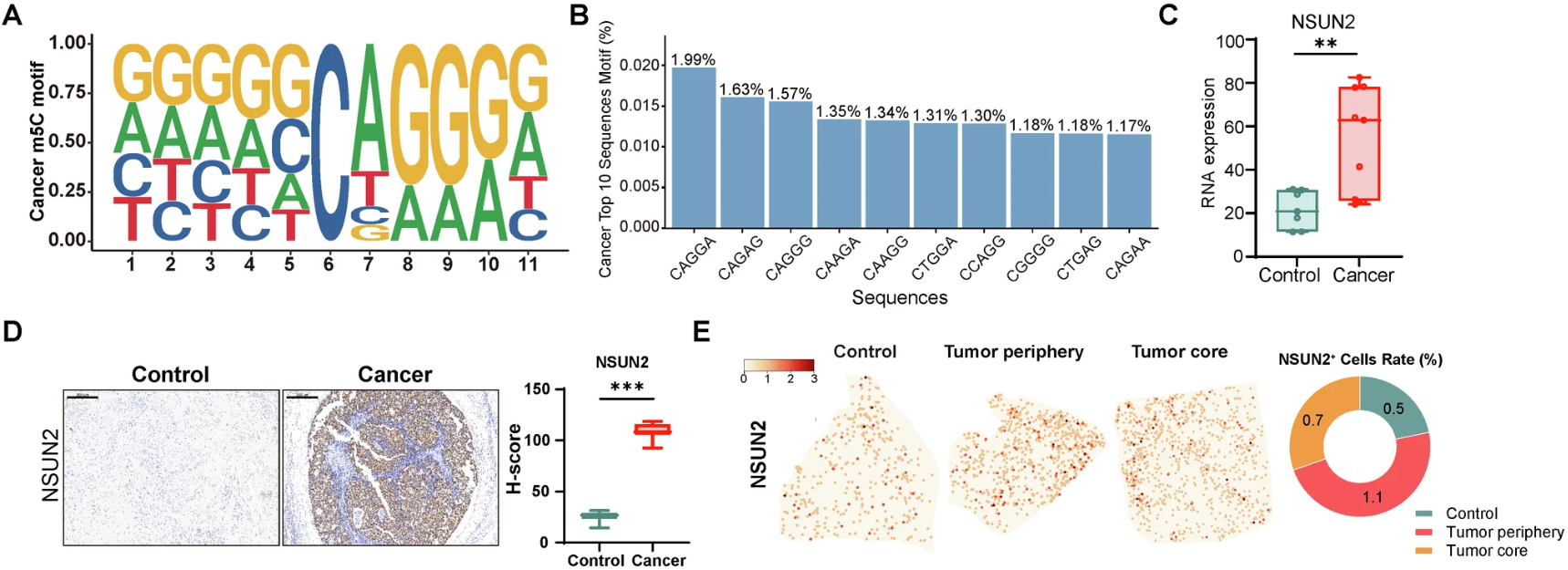
Figure 3. Molecular and spatial profiling of the NSUN2-m5C landscape. (A-B) Enriched sequence motifs and top sequence contexts for m5C sites identified in cervical cancer. (C-D) RNA-seq and IHC validation confirming significant NSUN2 upregulation at both transcript and protein levels in cancer versus control tissues. (E) High-definition Stereo-seq spatial mapping revealing the precise distribution and percentage of NSUN2+ cells across the tumor core and invasive periphery.
From a second perspective focused on chemotherapy resistance–associated expression, Stereo-seq further enabled the identification of functionally distinct resistant cell populations. Spatially resolved data showed that SERPINB5⁺ cells upregulate mitotic regulators such as CENPE, CDK1, and CCNB1, forming a transcriptional program that allows these cells to bypass chemotherapy-induced mitotic arrest. By accurately pinpointing where these resistant populations reside, Stereo-seq connects gene expression patterns with functional outcomes in situ. Without this level of spatial resolution, such subpopulations would be masked by signal averaging, making it difficult to link molecular mechanisms to therapeutic resistance. By overcoming the traditional trade-off between resolution and coverage, Stereo-seq converts fragmented transcriptomic signals into coherent spatial maps, enabling a clearer understanding of how the NSUN2–SERPINB5 axis drives tumor progression and drug resistance in a spatially dependent manner.
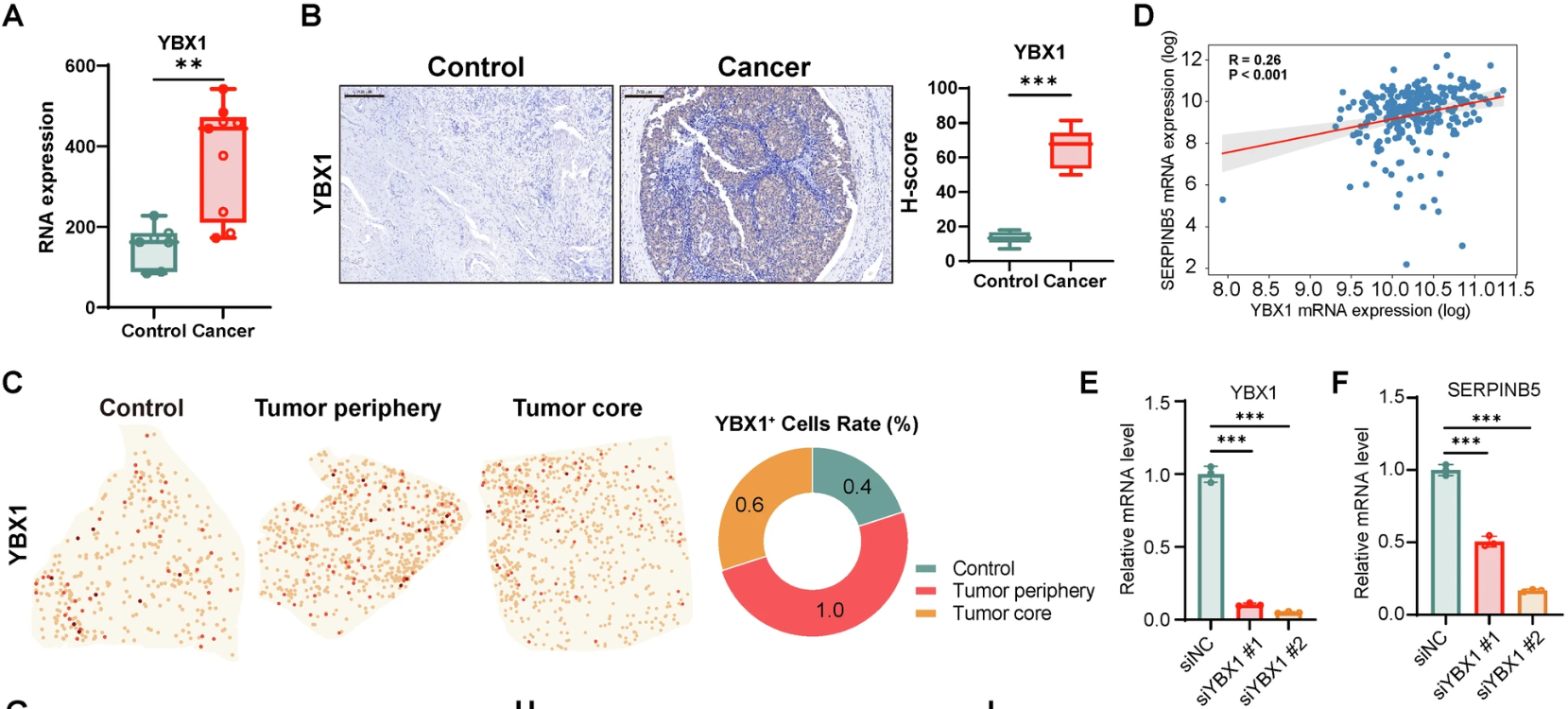
Figure 4. Validation of YBX1 as a spatial reader for the SERPINB5 axis. (A-B) Multi-level validation confirming significant YBX1 upregulation in cervical cancer at both transcript and protein levels. (C) Stereo-seq spatial mapping illustrating the distinct distribution of YBX1+ cells within the tumor core and periphery. (D-F) Correlation analysis and functional knockdown assays establishing the regulatory dependency of SERPINB5 expression on the m5C reader protein YBX1.
STOmics' Stereo-seq Portfolio for High-Resolution Mapping
Studies of epitranscriptomic regulation in cervical cancer—such as the spatially restricted NSUN2–SERPINB5 axis—highlight the need for technologies that can resolve molecular mechanisms within intact tissue context. To address this, STOmics offers a portfolio of spatiotemporal omics solutions powered by the Stereo-seq platform, enabling transcriptomic mapping at 500 nm resolution across centimeter-scale tissue regions, and supporting precise analysis of tumor heterogeneity and drug resistance.
Table 1. STOmics product matrix
Looking Ahead: Toward Integrated Spatial Multi-Omics
As cancer research advances toward precision medicine, the need for integrated, high-resolution data is rapidly increasing. Stereo-seq stands at the forefront of this transformation, enabling not only spatial transcriptomic mapping but also expansion into multi-omics through solutions such as spatial proteo-transcriptomics. Looking ahead, combining RNA modification profiling, spatial gene expression, and protein-level validation within the same tissue context will provide a more complete view of tumor biology, accelerate biomarker discovery, and support the development of more effective therapeutic strategies.
To explore how our spatiotemporal omics solutions can support your research in oncology or clinical immunology, please feel free to Contact Us or reach out via email at info_global@stomics.tech.
FAQs
Q: Why is m5C modification important in cancer research?
A: m5C is an important epitranscriptomic modification that regulates mRNA stability and translation, directly impacting oncogene expression such as SERPINB5. Dysregulation of m5C regulators like NSUN2 and YBX1 can enhance tumor growth, metastasis, and drug resistance, providing a critical regulatory layer beyond genetic mutations. Beyond cervical cancer, dysregulation of m5C writers such as NSUN2 has been reported across multiple cancer types, suggesting a broad oncogenic role for this epitranscriptomic layer."
Q: How does spatial information improve understanding of drug resistance?
A: Spatial analysis reveals not only the presence of resistant cells—such as SERPINB5⁺ populations—but also their precise location within the tumor. This helps uncover how these cells interact with the surrounding microenvironment and form localized resistance niches, which cannot be resolved by bulk or single-cell sequencing alone.
Q: What is the future direction of this technology?
A: The field is moving toward spatial multi-omics, integrating transcriptomic, proteomic, and epigenetic data within the same tissue context. This approach will enable deeper insights into disease mechanisms, improve biomarker discovery, and support more accurate and effective precision medicine strategies.
